Juan F. Poyatos
Group Leader
Research summary
Signals continuously impinge on cells, modifying their behaviour. This can be understood at three levels. At a population level, signals could modify the distribution of cell classes in a population, influencing core cell processes such as cell renewal in stem cell niches. At the intracellular pathway level, signals are effectively sensed and processed by combinations of genetic circuits. An open question is how these circuits distinguish signal from noise, and how circuit structure might limit this ability.
Publications
Rodrigo G, Bajic D, Elola I, Poyatos JF. Antagonistic autoregulation speeds up a homogeneous response in Escherichia coli. Sci Rep 2016; 6: 36196
Rodrigo G, Poyatos JF. Genetic Redundancies Enhance Information Transfer in Noisy Regulatory Circuits. PLoS Comput Biol 2016; 12: e1005156
Rodrigo G, Majer E, Prakash S, Daròs JA, Jaramillo A, Poyatos JF. Exploring the Dynamics and Mutational Landscape of Riboregulation with a Minimal Synthetic Circuit in Living Cells. Biophys J 2015; 109: 1070-6
Rodrigo G, Bajic D, Elola I, Poyatos JF. Dual autogenous control of the multiple antibiotic resistance phenotype in Escherichia coli. bioRxiv 2014; doi: 10.1101/008169
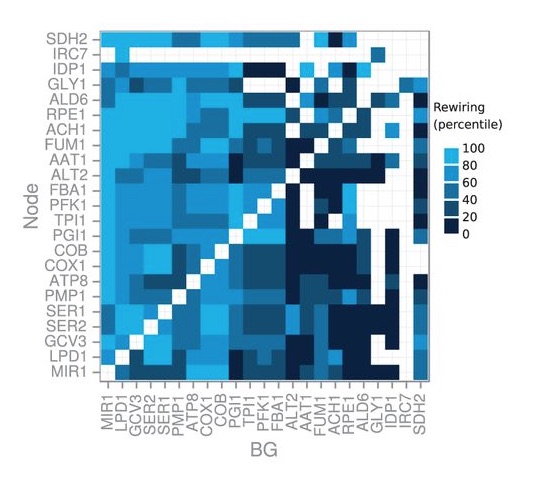
Bajic D, Moreno C, Poyatos JF. Rewiring of genetic networks in response to modification of genetic background. Genome Biol Evol 2014; 6:3267
 Two main problems were the centre of our research over the last two years. The first refers broadly to the limits of our present representation of biological systems as circuits and networks. The second problem deals with how the behaviour of individual cells determines the collective dynamics of cell populations.
Two main problems were the centre of our research over the last two years. The first refers broadly to the limits of our present representation of biological systems as circuits and networks. The second problem deals with how the behaviour of individual cells determines the collective dynamics of cell populations.
The representation of biological complexity.
We first analysed the stability of genetic networks. These networks help to map the molecular circuitry within cells by establishing genetic interactions between proteins (two proteins are “genetically” connected when the phenotype of the double mutant differs from that predicted for the combination of single mutations alone). The architecture of genetic networks can be modified with variation of genetic context (other mutations), which we studied by in silico modelling and by analysis of high-throughput data. Stability appeared to be closely connected to the mechanistic causes of robustness in the system under study. The second question refers to the assembly of transcriptional networks from its basic building blocks, or network motifs. We examined how the combination of functionally conflicting motifs could lead to trade-offs, and how such trade-offs can be resolved. We combined quantitative experiments and mathematical modelling in a bacterial model system with antagonistic autogenous control, the multiple antibiotic resistance regulatory module in Escherichia coli.
Individual behaviour and collective dynamics of cells.
Cells commonly cooperate among themselves by the secretion of molecules used by other cells (“public goods”). These interactions help establish cellular communities that are naturally unstable due to the rise of mutant individuals. These individuals avoid the costs associated to production of the public good, which can collapse the community. We analysed the persistence of these systems by examining a situation in which production of the public good by individual cells followed different strategies –constitutive, stochastic or plastic. We developed a mathematical model to quantify how each strategy would function in the presence of the non-contributing mutants, in a model that also controlled for a number of structural parameters. This analysis currently serves for the design and construction of robust bacterial communities in the lab.