Pablo Gastaminza Landart
Group Leader
Research summary
Our laboratory is interested in deciphering the molecular events underlying the functional dependence of HCV infection on lipid and lipoprotein metabolism and how to exploit these pathways to inhibit HCV infection. We use a cell culture infection system based on the infectious molecular clone JFH-1 and the Huh-7 hepatoma cell line and derived subclones to evaluate the impact that genetic and pharmacological manipulation of cellular lipid metabolism has on HCV infection and pathogenesis.
Publications
Galindo I. Et al. Antiviral drugs targeting endosomal membrane proteins inhibit distant animal and human pathogenic viruses. Antiviral Res. 2020 Nov26:104990. doi: 10.1016/j.antiviral.2020.104990.
Marcos-Villar L, Nistal-Villan E, Zamarreño N, Garaigorta U, Gastaminza P, Nieto A. Interferon-ß Stimulation Elicited by the Influenza Virus Is Regulated by the Histone Methylase Dot1L through the RIG-I-TRIM25 Signaling Axis. Cells. 2020 Mar 16;9(3):732
Castro V, Calvo G, Ávila-Pérez G, Dreux M, Gastaminza P. Differential Roles of Lipin1 and Lipin2 in the Hepatitis C Virus Replication Cycle. Cells. 2019 Nov 18;8(11):1456
Castro V, Ávila-Pérez G, Mingorance L, Gastaminza P. A Cell Culture Model for Persistent HCV Infection. Methods Mol Biol. 2019;1911:157-168
Sepúlveda-Crespo D, Jiménez JL, Gómez R, De La Mata FJ, Majano PL, Muñoz-Fernández MÁ, Gastaminza P. Polyanionic carbosilane dendrimers prevent hepatitis C virus infection in cell culture. Nanomedicine 2017; 13: 49-58
Our laboratory studies pathogenic human viral infections, and focuses on understanding the molecular basis of viral pathogenesis and identifying new molecular targets for antiviral therapy. Our final aim is to propose new therapeutic approaches for antiviral treatment and for reversion of virus-induced pathogenesis. To achieve these general aims, we have implemented cell culture models for infection by hepatitis C virus and other members of the Flaviviridae family such as dengue, Zika and West Nile viruses. Given the current health emergency due the COVID-19 pandemic, we have also implemented cell culture models of infection by SARS-CoV-2 coronavirus, including a compound screening platform for antiviral drug discovery.
Although their origin, nature and structure are not identical, a common feature of the aforementioned positive-strand RNA viruses is their ability to subvert host lipids and intracellular membranes to generate replication and assembly complexes. We previously reported that lipin1, a cellular enzyme that converts phosphatidic acid into diacylglycerol, is involved in the formation of the membranous web that hosts hepatitis C virus (HCV) replicase. In the liver, lipin1 cooperates with lipin2 to maintain glycerolipid homeostasis. We extended our previous study of the lipin family on HCV infection, by determining the impact of the lipin2 silencing on viral replication. Our data reveal that lipin2 silencing interferes with HCV virion secretion at late stages of the infection, without significantly affecting viral replication or assembly. Moreover, uninfected lipin2-, but not lipin1-deficient cells display alterations in mitochondrial and Golgi apparatus morphology, suggesting that lipin2 contributes to the maintenance of the overall organelle architecture. Finally, our data suggest a broader function of lipin2 for replication of HCV and other RNA viruses, in contrast with the specific impact of lipin1 silencing on HCV replication. Overall, our studies reveal distinctive functions of lipin1 and lipin2 in cells of hepatic origin, a context in which they are often considered functionally redundant.
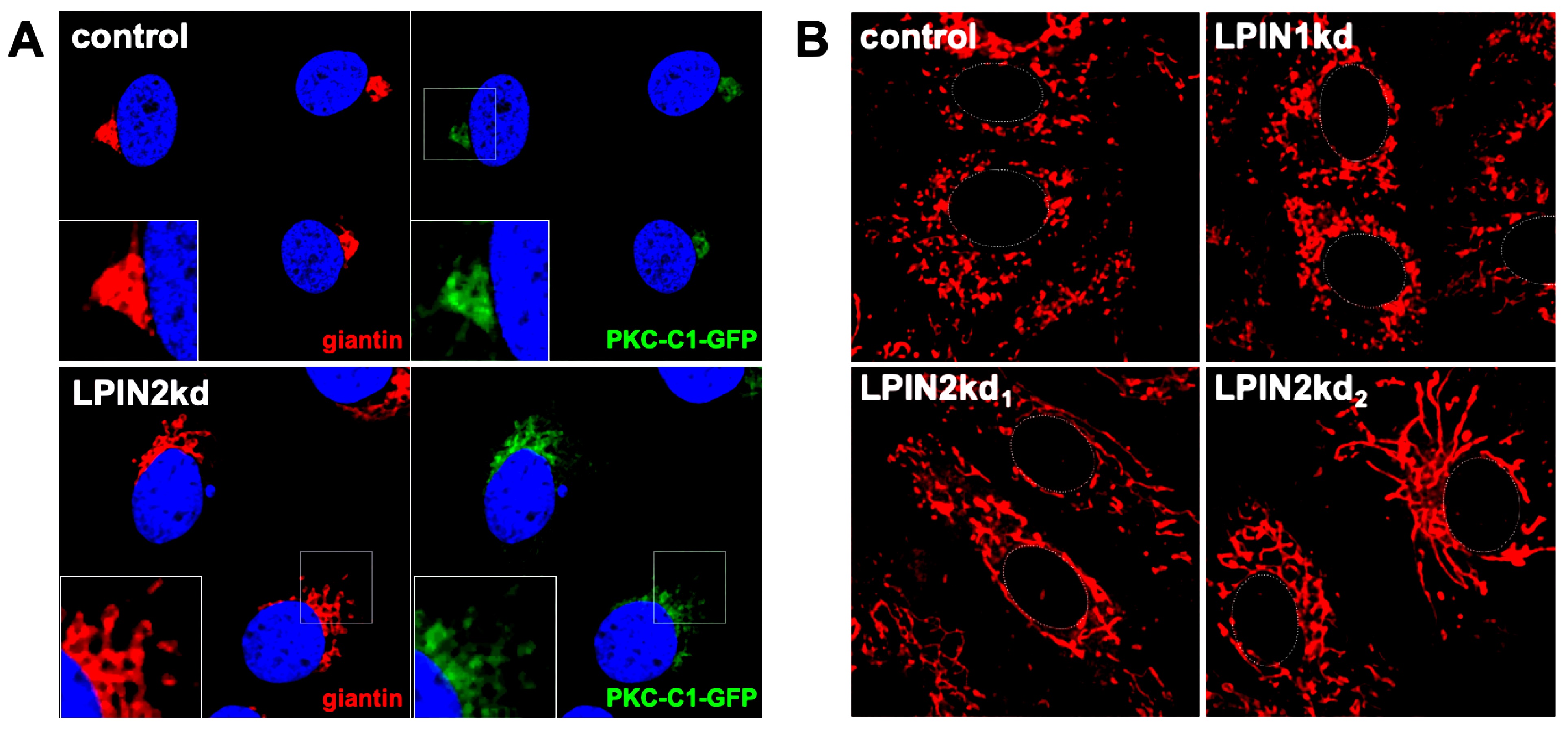
Lipin2, but not lipin1, silencing causes morphological alterations of the Golgi apparatus and mitochondrial elongation. Huh-7 cells constitutively expressing a DAG sensing probe (PKC-C1-D1-GFP) (see Materials and methods) were transduced with lentiviral vectors expressing non-targeting (control), shRNAs targeting LPIN1 (LPIN1kd) or LPIN2 mRNA (LPIN2kd1 and LPIN2kd2). At day 7 post transduction, control and lipin-deficient cultures expressing the DAG probe were fixed and processed for immunofluorescence microscopy using antibodies against a Golgi Apparatus marker (giantin) or stained with MitotrackerTM red following manufacturer recommendations (Life Technologies and imaged in vivo under a confocal microscope at 37oC and 5% CO2 to visualize mitochondria). A-Representative images of the Golgi morphology (red) and subcellular DAG probe (green) PKC-C1-GFP different cell lines. Nuclei were stained with DAPI and are shown in blue. were. B-Representative images of the mitochondrial morphology in the different cell populations. Cell nucleus is approximately delimited by a dotted white line for reference.








