Rafael Giraldo
Group Leader

Amyloids are stable protein assemblies that regulate phenotypes and enable their epigenetic inheritance. However, when they are the outcome of protein misfolding, amyloids can trigger diseases (i.e., human neurodegenerative and systemic amyloidosis). We create, through bottom-up Synthetic Biology, bio-resources based on bacterial amyloids with two major aims: i) understanding the molecular determinants of the shift between function and toxicity in natural amyloids; and ii) generating new devices based on amyloids as constructive resources in Biotechnology and Biomedicine.
Publications
Vendrell-Fernández S, Lozano-Picazo P, Cuadros-Sánchez P, Tejero-Ojeda MM, Giraldo R. Conversion of the OmpF porin into a device to gather amyloids on the E. coli outer membrane. ACS Synth Biol 2022; 11:655-667
Pantoja-Uceda D, Oroz J, Fernández C, de Alba E, Giraldo R, Laurents DV. Conformational priming of RepA-WH1 for functional amyloid conversion detected by NMR spectroscopy. Structure 2020; 28: 336-47.
Revilla-García A, Fernández C, Moreno-del Álamo M, de los Ríos V, Vorberg IM, Giraldo R. Intercellular transmission of a synthetic bacterial cytotoxic prion-like protein in mammalian cells. mBio 2020; 11: e02937-19.
Giraldo R. SynBio and the boundaries between functional and pathogenic RepA-WH1 bacterial amyloids. mSystems 2020; 5: e00553-20.
Giraldo R. Optogenetic navigation of routes leading to protein amyloidogenesis in bacteria . J Mol Biol 2019; doi: 10.1016/j.jmb.2019.01.037
Fernández C, Giraldo R Modulation of the aggregation of the prion-like protein RepA-WH1 by chaperones in a cell-free expression system and in cytomimetic lipid vesicles. ACS Synth Biol 2018; 7: 2087-2093
RepA is a protein essential for the maintenance of plasmids in Gram-negative bacteria whose WH1 domain undergoes conformational changes capacitating it either as a transcriptional repressor, as DNA replication initiator or, through its assembly into amyloid oligomers, to hinder premature re-replication rounds. Stable RepA-WH1 dimers become metastable amyloidogenic monomers upon allosteric binding of plasmid-specific dsDNA sequences, or acidic phospholipids. We have engineered fusions of mutant variants of RepA-WH1 to the fluorescent protein mCherry, generating a prion-like protein (prionoid) that is epigenetically transmitted from mother-to-daughter Escherichia coli cells under the modulation of the Hsp70 chaperone DnaK, causing a synthetic ‘generic’ amyloid proteinopathy. In the bacterial cytosol, RepA-WH1 aggregates as distinct strains, either dense foci or fluidized particles, which exhibit acute and mild cytotoxicity, respectively. RepA-WH1 amyloidosis recapitulates in bacteria the mitochondrial route in human amyloid diseases, including the formation of oligomeric pores at the internal membrane, the generation of reactive oxygen species and the co-aggregation of stress-responsive factors. We have demonstrated that RepA-WH1 can be horizontally transmitted to mammalian cultured cells, but its cytotoxicity depends on its heterologous expression in the recipient cells (Fig. 1A).
We have used RepA-WH1 as a benchmark for the design of synthetic tools to probe protein amyloidogenesis, including gold nanoparticles-based sensors and screening devices exploiting amyloid-promoted overriding of translation termination, both in yeast and in bacteria. We have also achieved control on RepA-WH1 amyloidogenesis through optogenetics, i.e., the fusion of a blue light-responsive plant domain (LOV2) to WH1. Under illumination, a LOV2-WH1-mCherry fusion assembles as oligomers that are cytotoxic in E. coli, hampering bacterial growth (as novel antimicrobial ‘optobiotics’), while in the darkness it forms inert amyloid fibrils and hydrogels (Fig. 1B).
Our latest development is engineering the E. coli outer membrane porin OmpF by grafting an amyloidogenic sequence in its extracellular loop L5, thus enticing to the bacterial envelope, through the assembly of amyloids, the same sequence when presented as a free peptide or in its protein of origin. Targeting bacteria to amyloids opens a way to the bioremediation of prions persistent in natural environments (Fig. 1C).
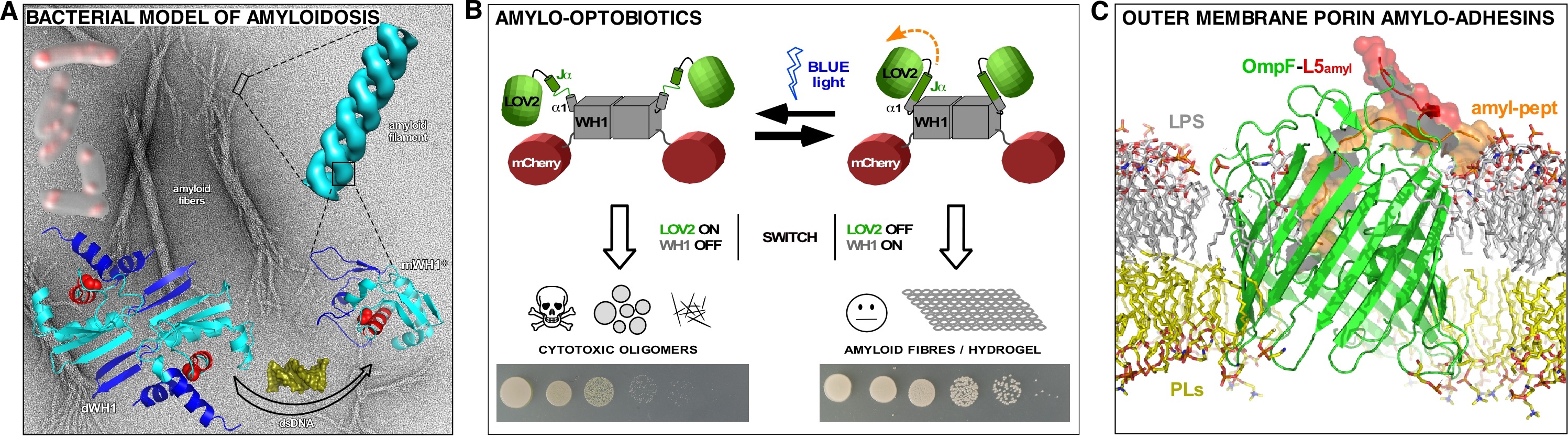
Figure legend. A bacterial protein ‘moonlighting’ as a functional amyloid (RepA-WH1, panels A and B) can be either engineered as a minimal model of amyloid disease (A), or tailored to modulate amyloid toxicity with light constituting a potential antimicrobial agent (B). Bacterial outer membrane porins (OmpF, panel C) can be converted into scavengers of environmental amyloids.
.







