Jesús Blazquez
Group Leader
Research summary
We try to understand the genetic mechanisms involved in genome stability in bacteria and their roles on evolution and adaptation. Specifically, we study both stable and induced hypermutation/hyper-recombination as “bacterial strategies” to speed adaptation to environmental stress. This knowledge may be useful to i) prevent the development of antibiotic resistance in bacterial pathogens (e.g. Escherichia coli, Pseudomonas aeruginosa, Mycobacterium tuberculosis, etc) and ii) to improve prokaryotic species of industrial interest (e.g. Streptomyces, Bifidobacterium, Rhodococcus, Corynebacterium, Pyrococcus, Halobacterium, etc).
Publications
Castañeda-García A, Martín-Blecua I, Cebrián-Sastre E, Chiner-Oms A, Torres-Puente M, Comas I, and Blázquez J. Specificity and mutagenesis bias of the mycobacterial alternative mismatch repair analyzed by mutation accumulation studies. Sci Adv 2020; 6, eaay4453. DOI: 10.1126/sciadv.aay4453.
Rodriguez-Rosado AI, Valencia EI, Costas C, Galhardo RS, Rodriguez-Beltan J, and Blazquez, J N-acetylcisteine blocks SOS induction and mutagenesis produced by fluoroquinolones in Escherichia coli. J. Antimicrob. Chemother 2019; 74: 2188-2196. doi: 10.1093/jac/dkz210
Blazquez J, Rodríguez-Beltrán J, and Matic IAntibiotic-induced genetic variation: How it arises and how it can be prevented. Ann. Rev. Microbiol 2018; 72:209-230. doi: 10.1146/annurev-micro-090817-062139
Castañeda-García A, Prieto AI, Rodríguez-Beltrán J, Alonso N, Cantillon D, Costas C, Pérez L, Zegeye ED, Herranz M, Plociński P, Tonjum T, García de Viedma D, Paget M, Waddell SJ, Rojas AM, Doherty AJ and Blázquez J. A non-canonical mismatch repair pathway in Prokaryotes. Nat Comm 2017; 8:14246
Martín-Gutiérrez G, Rodríguez-Beltrán J, Rodríguez-Martínez JM, Costas C, Aznar J, Pascual Á, Blázquez J. Urinary tract physiological conditions promote ciprofloxacin resistance in low-level-quinolone-resistant Escherichia coli. Antimicrob Agents Chemother 2016; 60: 4252-8
We try to understand the genetic mechanisms involved in genome stability in bacteria and their roles on evolution and adaptation. Specifically, we study both stable and induced hypermutation/hyper-recombination as “bacterial strategies” to speed adaptation to environmental stress. This knowledge may be useful to i) prevent the development of antibiotic resistance in bacterial pathogens (e.g. Escherichia coli, Pseudomonas aeruginosa, M. tuberculosis, etc) and ii) to improve prokaryotic species of industrial interest (e.g. Streptomyces, Bifidobacterium, Rhodococcus, Corynebacterium, Pyrococcus, Halobacterium, etc).
Main research lines:
1. Regulation of hypermutation and hyper-recombination.
· Transcriptional regulation of specialised DNA polymerases (SOS regulon)
· Regulation of mismatch repair
· Effect of antibiotics on bacterial mutation and recombination: antibiotics as promoters of antibiotic resistance?
2. Genetic basis of genome stability in prokaryotes lacking a canonical MutS-MutL-based DNA mismatch repair (MMR) system: we are studying the possible existence of alternative MMR pathways in these organisms, including most Actinobacteria (e.g. Mycobacterium, Streptomyces, Rhodococcus, etc) and some Archaea (Pyrococcus, Halobacterium, etc).
3. Development of hypermutant/hyper-recombinant prokaryotes of industrial interest as biotechnological tools to produce improved biosynthetic or degrading pathways. Also, we explore the possibility of developing genetically superstable strains.
4. Preventing emergence and evolution of antibiotic resistance: search and characterize compounds able to inhibit the development of antibiotic resistance (by mutation, recombination and horizontal transfer).
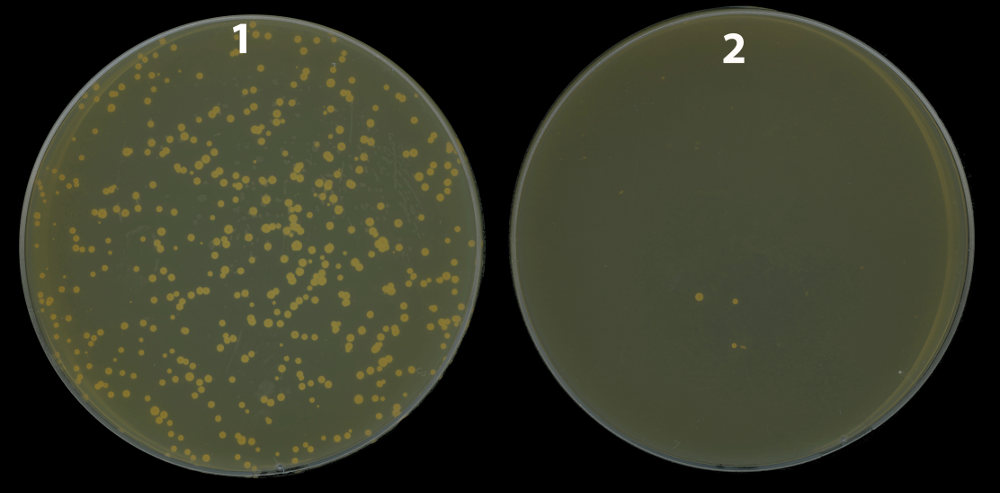
A strain of Mycobacterium in which nucS gene has been eliminated (1) produces a high number of mutants resistant to the antibiotic present in the Petri dish (rifampicin). However, the parental strain (with nucS active) produces much less mutants (2)./ Alfredo Castañeda and Jesús Blázquez, CNB-CSIC.








