Pilar Cubas
Group Leader
Research summary
We are studying the genetic basis of the control of axillary bud development in the model system Arabidopsis, and in the crop species tomato and potato in which control of lateral shoot branching is of great agronomical interest. We have characterised the Arabidopsis BRANCHED1 (BRC1) gene, which acts as a central switch of axillary bud development and outgrowth. We are now expanding our knowledge of the genetic networks involving BRC1 in Arabidopsis.
Publications
Nicolas M, Torres-Pérez R, Wahl V, Cruz-Oró E, Rodríguez-Buey ML, Zamarreño AM, Martín-Jouve B, García-Mina JM, Oliveros JC, Prat S & Cubas P. Spatial control of potato tuberisation by the TCP transcription factor BRANCHED1b. Nature Plants. 2022 8, 281–294.
Li Q, Sánchez Martín-Fontecha E, Khosla A, White A, Chang S, Cubas P and Nelson D. Degradation of SUPPRESSOR OF MAX2 1 (SMAX1) by the strigolactone receptor D14 in Arabidopsis thaliana. Plant communications. 2022, 3, (2)10030.
Fichtner F, Barbier F, Kerr S, Dudley C, Cubas P, Turnbull C, Brewer P, Beveridge C. Different plasticity of bud outgrowth at cauline and rosette nodes in Arabidopsis thaliana Plant Physiology 2021.
Yang Y, Nicolas M, Zhang J, Yu H, Guo D, Yuan R, Zhang T, Yang J, Cubas P, Qin G. The TIE1 transcriptional repressor controls shoot branching by directly repressing BRANCHED1 in Arabidopsis. PLoS Genet. 2018 Mar 23;14(3):e1007296.
González-Grandío E, Pajoro A, Franco-Zorrilla JM, Tarancón C, Immink RG, Cubas P. Abscisic acid signaling is controlled by a BRANCHED1/HD-ZIP I cascade in Arabidopsis axillary buds. Proc Natl Acad Sci U S A. 2017; 114(2):E245-E254
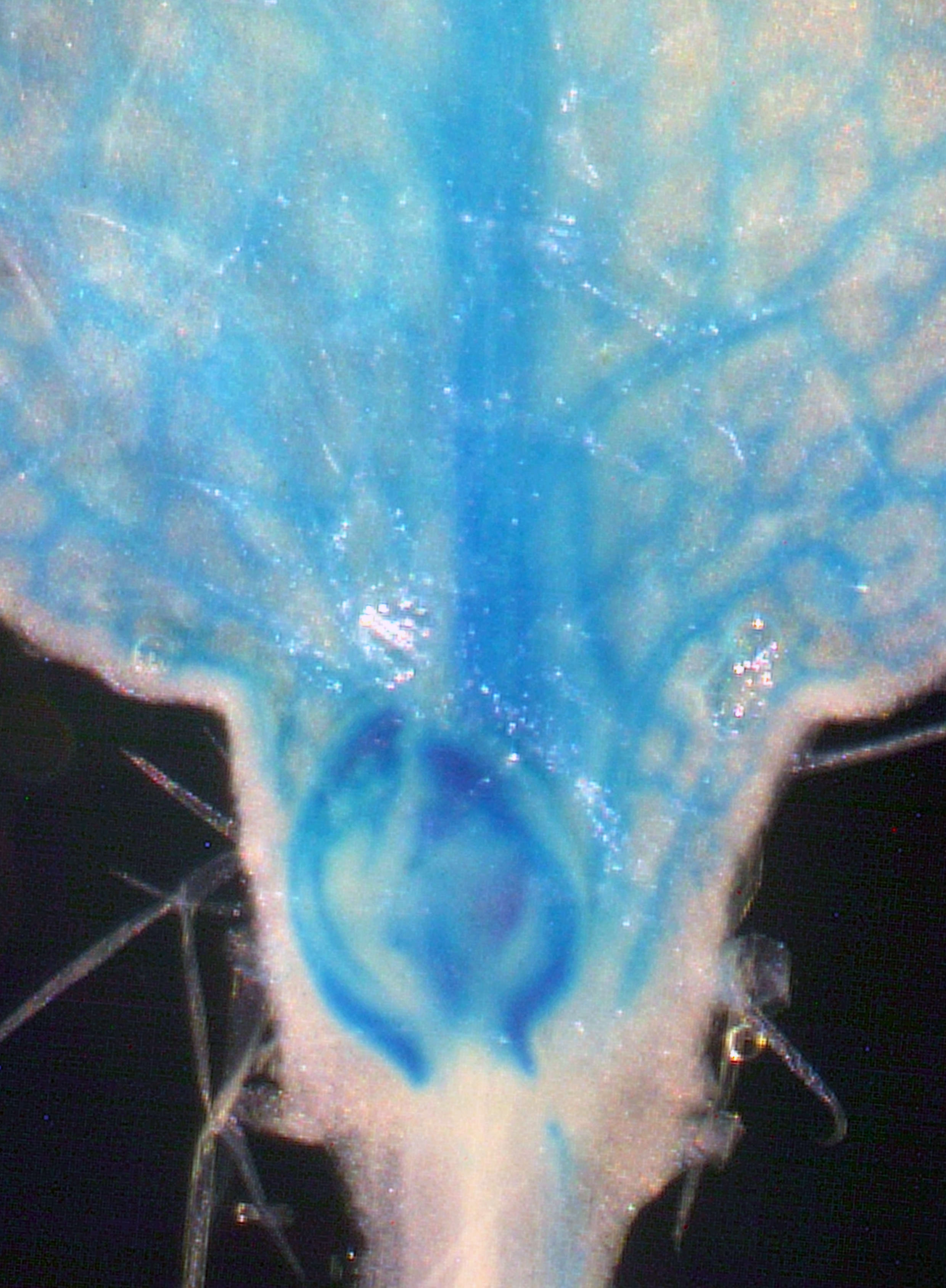 Shoot branching patterns depend on a key developmental decision: whether axillary buds grow out to give a branch or remain dormant in the leaf axils. This decision is controlled by hormone-mediated endogenous and environmental stimuli. A decrease in the red to far-red light ratio (R:FR) –a sign of shading by neighbouring vegetation– triggers a set of developmental plant responses termed shade avoidance syndrome. One of these responses is suppression of axillary bud outgrowth.
Shoot branching patterns depend on a key developmental decision: whether axillary buds grow out to give a branch or remain dormant in the leaf axils. This decision is controlled by hormone-mediated endogenous and environmental stimuli. A decrease in the red to far-red light ratio (R:FR) –a sign of shading by neighbouring vegetation– triggers a set of developmental plant responses termed shade avoidance syndrome. One of these responses is suppression of axillary bud outgrowth.
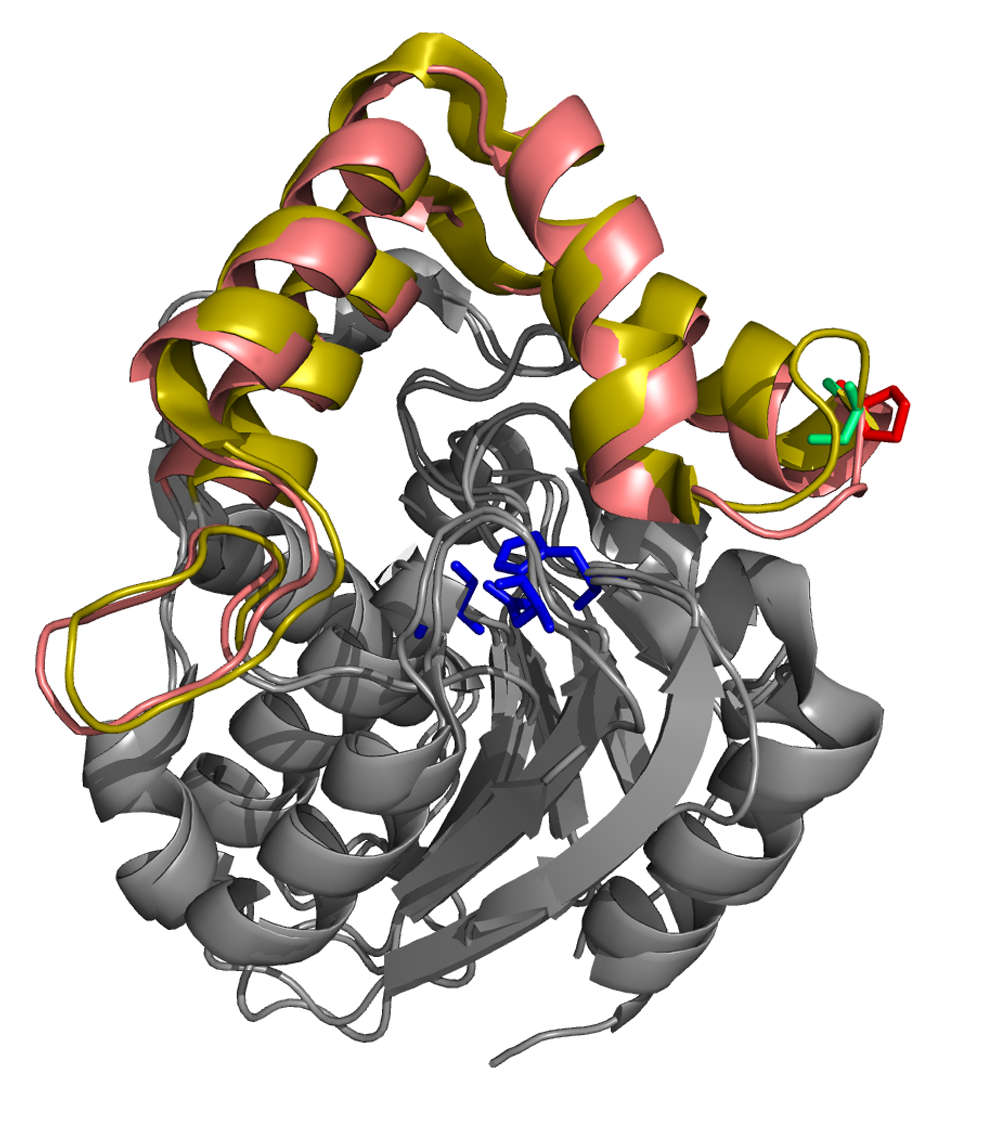
The Arabidopsis gene BRANCHED1, which encodes a TCP transcription factor, is a point at which signals that suppress shoot branching are integrated in axillary buds. We showed that BRC1 is necessary for branch suppression in response to shade, and its BRC1 transcription is positively regulated after exposure to low R:FR.
To understand the growth-to-dormancy transition in axillary buds, we compared transcriptomic profiles of wild-type and brc1 axillary buds and identified sets of genes that might be controlled directly by BRC1. We distinguished a set of upregulate abscisic acid response genes and two networks of cell cycle- and ribosome-related downregulated genes. The downregulated genes have promoters enriched in TCP binding sites, which suggests transcriptional regulation by TCP factors. We compared our transcriptomic data with two additional “active vs dormant bud” transcriptomic data sets and found “core” coregulated gene networks closely associated to each condition.
Strigolactones (SL) are phytohormones that regulate shoot branching. SL perception and signalling involves the F-box protein MAX2 and the hydrolase D14, proposed to act as a SL receptor. We used strong loss-of-function alleles of the D14 gene to characterise its function. Our data showed that D14 distribution overlaps that of MAX2 at tissue and subcellular levels, allowing physical interactions between these proteins. Grafting studies indicated that neither D14 mRNA nor the protein move upwards over a long range in the host. Like MAX2, D14 is needed locally in the aerial part of the plant to suppress shoot branching. We also identified a mechanism of SL-induced MAX2-dependent proteasome-mediated D14 degradation. This negative feedback loop wouldcause a substantial drop in SL perception, which would effectively limit SL duration and signalling intensity.