Jaime Martín Benito
Group Leader
The main research line of our group is the study of the influenza A ribonucleoproteins (RNPs) that conform the virus nucleocapsid.
Publications
Coloma R, Arranz R, de la Rosa-Trevin JM, Sorzano COS, Munier S, Carlero D, Naffakh N, Ortín J, Martin-Benito J.The processive helical track: A structural model for influenza virus transcription and replication. Nat Microbiology 2020; 5: 727-734.Franco A, Arranz R, Fernández-Rivero N, Velázquez-Campoy A, Martín-Benito J, Segura J, Prado A, Valpuesta JM, Muga A. Structural insights into the ability of nucleoplasmin to assemble and chaperone histone octamers for DNA deposition. Scientific Reports. 2019 Jul 1;9(1):9487.
Quintana-Gallardo L, Martín-Benito J, Marcilla M, Espadas G, Sabidó E, Valpuesta JM. The cochaperone CHIP marks Hsp70- and Hsp90-bound substrates for degradation through a very flexible mechanism. Scientific Reports. 2019 Mar 25;9(1):5102.
Fernández-Justel D, Núñez R, Martín-Benito J, Jimeno D, González-López A, Soriano EM, Revuelta JL, Buey RM. A Nucleotide-Dependent Conformational Switch Controls the Polymerization of Human IMP Dehydrogenases to Modulate their Catalytic Activity. Journal of Molecular Biology. 2019 Mar 1;431(5):956-969.
Arranz R, Martín-Benito J, Valpuesta JM. Structure and Function of the Cochaperone Prefoldin. Adv Exp Med Biol 2018; 1106:119-131.
Arranz R, Coloma R, Chichón FJ, Conesa JJ, Carrascosa JL, Valpuesta JM, Ortín J, Martín-Benito J. The structure of native influenza virion ribonucleoproteins. Science 2012; 338(6114):1634-7.
The main line of the group is focused on the study of the Influenza A ribonucleoproteins (RNPs) that conforms the virus nucleocapsid. RNPs are macromolecular complexes composed of the genomic RNA bound to multiple monomers of a nucleoprotein and a single copy of the polymerase. In recent years our laboratory has determined the structure of the isolated RNPs at medium resolution and we have verified that this structure is present in native virions using cryoelectron tomography. Currently our research on this topic is following two major subjects that will extend to the next years.
The first one is the resolution improvement of the RNP structure, and for that we will make use of the state-of-the-art electron cryomicroscopes equipped with direct electron detectors, recently acquired by the CNB. We have already discovered the existence of an enormous conformational variability in the RNPs structure, and this has been possible through the design of a new protocol able to classify and reconstruct helical structures. We have demonstrated that this extreme conformational variability is closely related with the biological roles of the RNPs. With this idea we opened the second major line of our research, the elucidation of the structural basis of transcription and replication mechanisms and for that our plan is to complement structural data with biochemical assays that will allow us to establish the action mechanism. Our research also extends to the study of influenza virus polymerase at high resolution. Our goal is to solve the polymerase dimer structure that represents the functional complex for replication of RNPs.
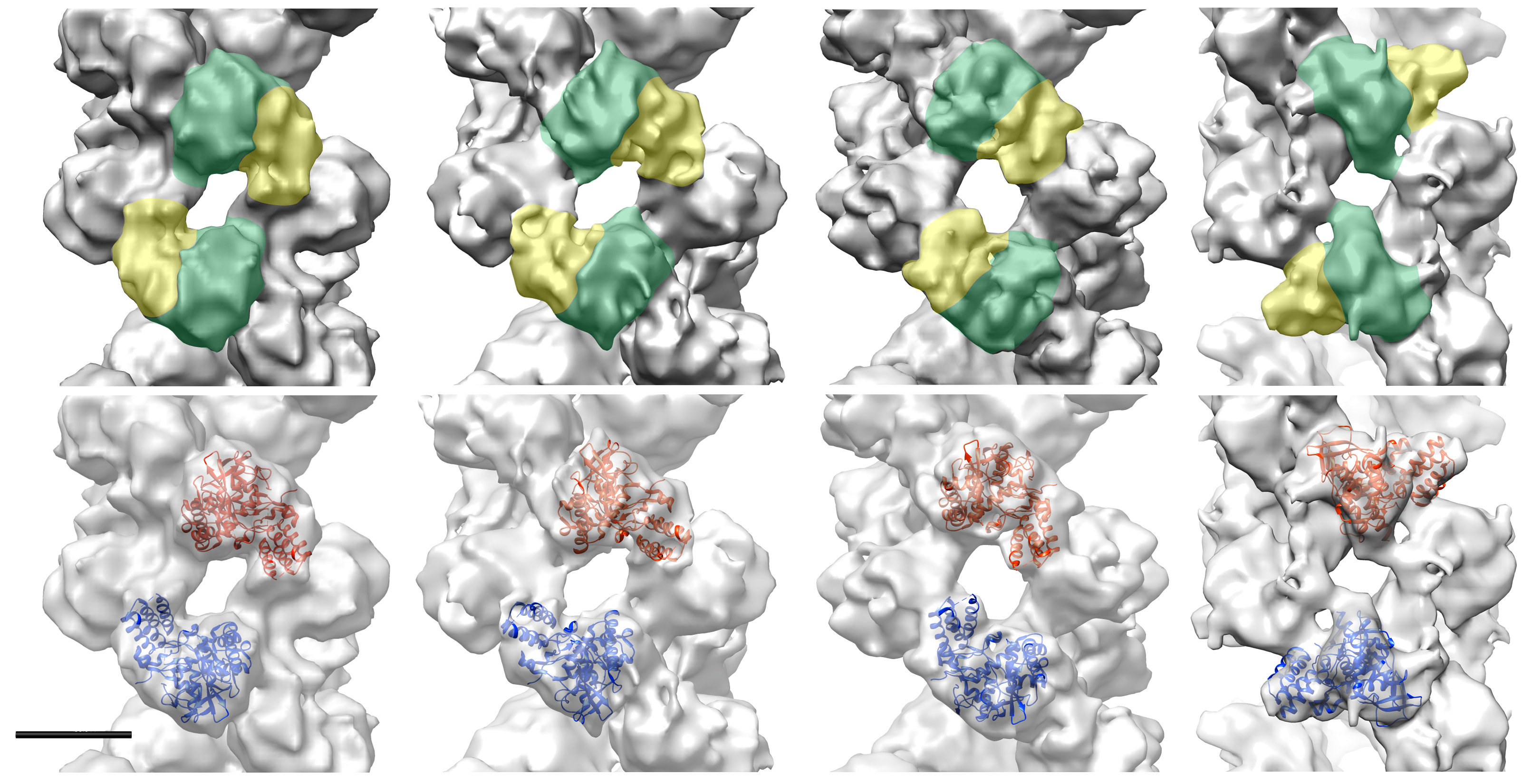 3D reconstructions of different conformations of the helical part of the influenza virus ribonucleoproteins. Top row: the head and body domains of the nucleoproteins are outlined in yellow and green, respectively. Bottom row: the docking of the influenza A nucleoprotein atomic structure (pdb 2IQH) is shown in the opposite strands in red and blue. Scale bar, 50Å.
3D reconstructions of different conformations of the helical part of the influenza virus ribonucleoproteins. Top row: the head and body domains of the nucleoproteins are outlined in yellow and green, respectively. Bottom row: the docking of the influenza A nucleoprotein atomic structure (pdb 2IQH) is shown in the opposite strands in red and blue. Scale bar, 50Å.
.







