Jose Manuel Franco
Group Leader
Research summary
Plant plasticity during adaptation to the environment involves specific transcriptional signal-response networks that allow them to reprogram their growth and development. We are interested in studying sequence-specific transcription factors (TFs), the regulatory proteins responsible for the transcriptional activation or repression of target genes.
Publications
Hajheidari M, Wang Y, Bhatia N, Vuolo F, Franco-Zorrilla JM, et al. Autoregulation of RCO by low-affinity binding modulates cytokinin action and shapes leaf diversity. Curr Biol 2019; 29: 4183-4192.e6.
Peñuelas M, Monte I, Schweizer F, Vallat A, Reymond P, García-Casado G, Franco-Zorrilla JM, Solano R. Jasmonate-Related MYC Transcription Factors Are Functionally Conserved in Marchantia polymorpha. The Plant Cell 2019; DOI: https://doi.org/10.1105/tpc.18.00974
Monte I, Franco-Zorrilla JM, García-Casado G, Angel M. Zamarreño, José M. García-Mina, Ryuichi Nishihama, Takayuki Kohchi and Roberto Solano. A Single JAZ Repressor Controls the Jasmonate Pathway in Marchantia polymorpha. Mol Plant. 2019;12(2):185‐198. doi:10.1016/j.molp.2018.12.017
Silva CS, Nayak A, Lai X, Hutin S, Hugouvieux V, et al. Molecular mechanisms of Evening Complex activity in Arabidopsis. Proc Natl Acad Sci USA 2020; 117: 6901-6909.
Ortigosa A, Fonseca S, Franco‐Zorrilla JM, Fernández‐Calvo P, Zander M, Lewsey MG, García‐Casado G, Fernández‐Barbero G, Ecker JR, Solano R. The JA‐pathway MYC transcription factors regulate photomorphogenic responses by targeting HY5 gene expression. The Plant Journal 2020 DOI: 10.1111/tpj.14618
Plant plasticity during adaptation to the environment involves specific transcriptional signal-response networks that allow them to reprogram their growth and development. Regulation of these networks relies on sequence-specific transcription factors (TFs), regulatory proteins responsible for the transcriptional activation or repression of target genes.
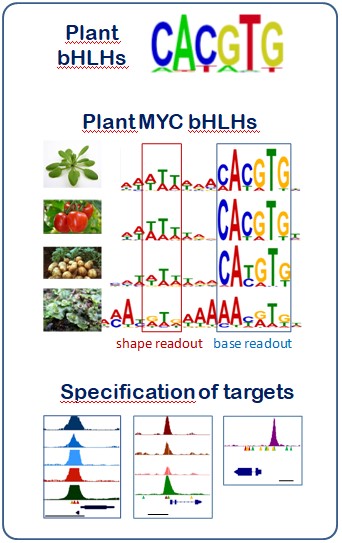
Research in our group is focused in the study of the components that determine specific recognition of TF target genes and which may influence in the levels of gene expression. During the last few years we have contributed to the characterization of one of these components, such as the short DNA sequences bound by TFs, known as TF-binding sites (TFBS). Despite TFBS sequence is the major factor determining target recognition, during the last two years we have explored the role of some other components involved in this process. With this regard, we have demonstrated that binding of some TFs extends beyond the TFBS core sequence, as some distant nucleotides, likely determining DNA-shape, are necessary for protein binding. We are also studying the role of the cytosine methylation epigenetic mark in the TFBS region during TF-target recognition, as well as its genetic control, what will allow adding a new layer of regulation of gene expression
In parallel to the experimental approaches, we are developing some easy-to-use bioinformatic tools useful for the interpretation of transcriptional data and for the prediction of TFBS involved in the regulation of biological processes. These tools would contribute to a better and faster interpretation of biological data for the plant biology community, particularly in the case of non-expert researchers in bioinformatics or in the study of non-model species.