José Luis Martínez
Group Leader
Research summary
Several infections at hospitals are due to opportunistic pathogens, microorganisms that rarely infect healthy people, but are a frequent cause of infection in people with basal diseases, who are immunodepressed or debilitated. Environmental bacteria, frequently antibiotic resistant, constitute a large percentage of those pathogens. Our work focuses on understanding the mechanisms of virulence and resistance, as well as possible crosstalk, of these pathogens.
Publications
Hernando-Amado S, Sanz-García F, Martínez JL. Rapid and robust evolution of collateral sensitivity in Pseudomonas aeruginosa antibiotic-resistant mutants. Sci Adv 2020; 6 (32): eaba5493.
Gil-Gil T, Corona F, Martínez JL, Bernardini A. The inactivation of enzymes belonging to the central carbon metabolism is a novel mechanism of developing antibiotic resistance. mSystems. 2020; 5 (3): e00282-20.
Hernando-Amado S, Sanz-Garcia F, Martinez JL. Antibiotic resistance evolution is contingent on the quorum sensing response in Pseudomonas aeruginosa. Mol Biol Evol. 2019; 36 (10): 2238–2251.
Hernando-Amado S, Coque TM, Baquero F, Martinez JL. Defining and combating antibiotic resistance from One Health and Global Health perspectives. Nat Microbiol 2019; 4 (9): 1432-1442.
Ruppe E, Ghozlane A, Tap J, Pons N, Alvarez AS, et al. Prediction of the intestinal resistome by a novel 3D-based method. Nat Microbiol 2019; 4: 112-123.
We study the biology of opportunistic pathogens, focusing on the networks and the evolutionary processes that connect resistance and virulence. In the last years, we have standardised some tools, based on experimental evolution, whole-genome sequencing and functional assays, for predicting the evolution of antibiotic resistance and the consequences of acquiring such resistance for bacterial physiology. Using these approaches, we characterised mechanisms of resistance to last-generation antibiotics and combination of them. Notably, acquisition of resistance is linked to changes in the susceptibility of other antibiotics besides those used for selection. In this regard, we have determined the networks of cross-resistance and collateral susceptibility associated to the acquisition of resistance to different antibiotics.
One important element in our studies is determining the elements that modulate the robustness and predictability of evolutionary trajectories towards antibiotic resistance of bacterial pathogens. Robustness is particularly relevant for exploiting the information concerning collateral susceptibility in order to implement more efficient therapeutic strategies based in antibiotic combinations or cycling. We have been working in this topic and have found some robust collateral susceptibility networks that will be explored in clinical strains in the next future. Among the elements that drive the evolution of antibiotic resistance from stochasticity to determinism, we are particularly interested in the epistatic interactions between elements involved in antibiotic resistance and virulence of bacterial pathogens. Besides mutation-driven resistance, we are studying compounds or conditions that can lead to transient resistance, as well as those that can increase the susceptibility to antibiotics and could hence be used as co-adjuvants in therapy.
A final aspect of our work concerns the One Health and Global Health aspects of antibiotic resistance. We contributed to the development of novel tools and metagenomic analyses for analysing the role of different non-clinical habitats in the evolution and spread of antibiotic resistance.
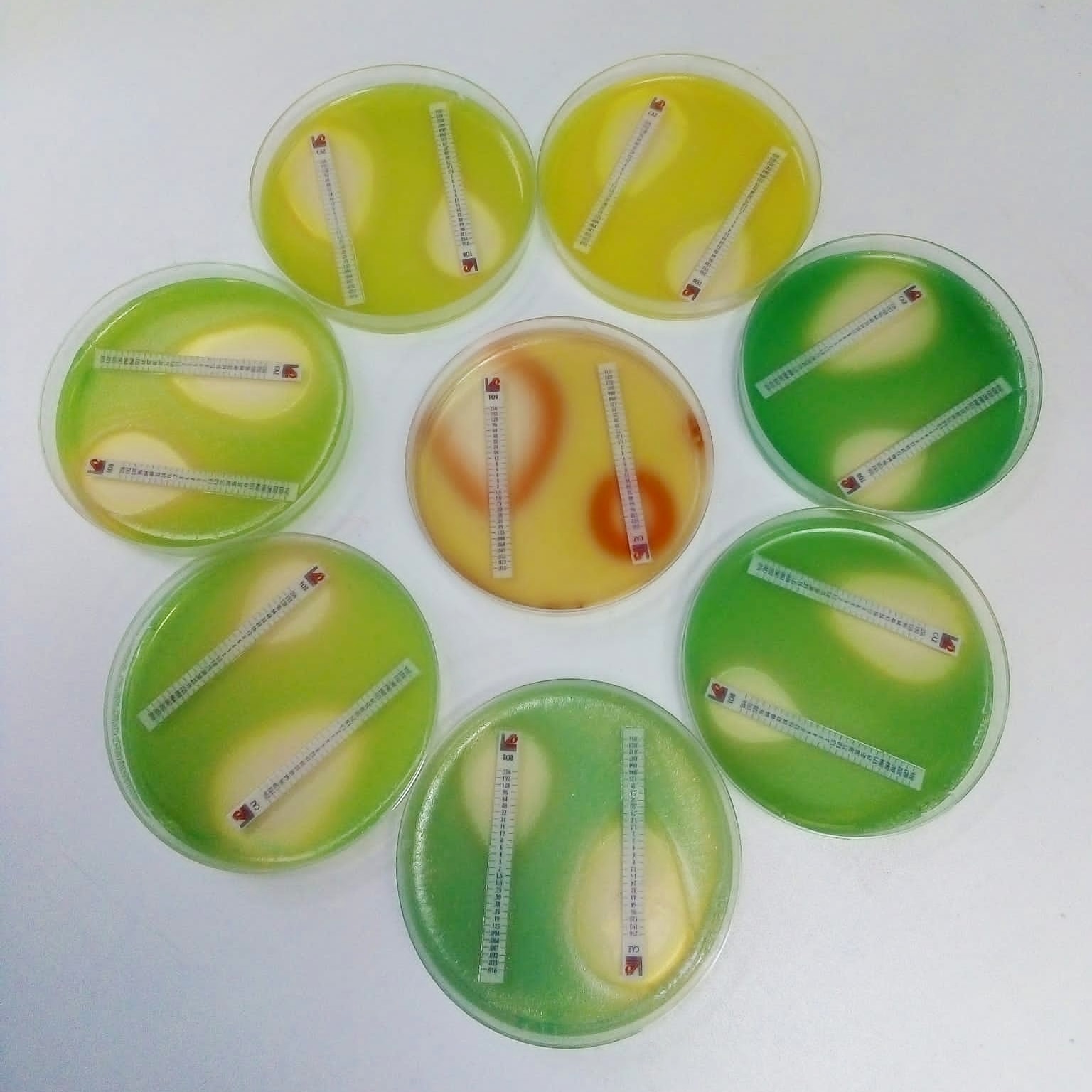
Figure Legend: Radiative evolution of antibiotic resistance in presence of antibiotics. Evolution is generally considered a very slow process. However, microorganisms, and bacteria in particular, reach a large number of generations in a very short period of time. This makes it possible to use Adaptive Laboratory Evolution (ALE) experiments to predict bacterial evolution in the presence of different selective pressures, such as antibiotics. This photograph shows the different changes in pigmentation and antibiotic resistance level (measured with E-Test strips), of the opportunistic pathogen Pseudomonas aeruginosa, after only 21 days of ALE in the presence of different concentrations of tobramycin.








