Infect Immun. 2012 Dec;80(12):4071-7
Garber JJ, Takeshima F, Anton IM, Oyoshi MK, Lyubimova A, Kapoor A, Shibata T, Chen F, Alt FW, Geha RS, Leong JM, Snapper SB.
 The human pathogens enteropathogenic Escherichia coli (EPEC) and vaccinia virus trigger actin assembly in host cells by activating the host adaptor Nck and the actin nucleation promoter neural Wiskott-Aldrich syndrome protein (N-WASP). EPEC translocates effector molecules into host cells via type III secretion, and the interaction between the translocated intimin receptor (Tir) and the bacterial membrane protein intimin stimulates Nck and N-WASP recruitment, leading to the formation of actin pedestals beneath adherent bacteria. Vaccinia virus also recruits Nck and N-WASP to generate actin tails that promote cell-to-cell spread of the virus.
The human pathogens enteropathogenic Escherichia coli (EPEC) and vaccinia virus trigger actin assembly in host cells by activating the host adaptor Nck and the actin nucleation promoter neural Wiskott-Aldrich syndrome protein (N-WASP). EPEC translocates effector molecules into host cells via type III secretion, and the interaction between the translocated intimin receptor (Tir) and the bacterial membrane protein intimin stimulates Nck and N-WASP recruitment, leading to the formation of actin pedestals beneath adherent bacteria. Vaccinia virus also recruits Nck and N-WASP to generate actin tails that promote cell-to-cell spread of the virus.
In addition to Nck and N-WASP, WASP-interacting protein (WIP) localizes to vaccinia virus tails, and inhibition of actin tail formation upon ectopic expression of WIP mutants led to the suggestion that WIP is required for this process. Similar studies of WIP mutants, however, did not affect the ability of EPEC to form actin pedestals, arguing against an essential role for WIP in EPEC-induced actin assembly. In this study, we demonstrate that Nck and N-WASP are normally recruited by vaccinia virus and EPEC in the absence of WIP, and neither WIP nor the WIP family members CR16 and WIRE/WICH are essential for pathogen induced actin assembly. In addition, although Nck binds EPEC Tir directly, N-WASP is required for its localization during pedestal formation.
Overall, these data highlight similar pathogenic strategies shared by EPEC and vaccinia virus by demonstrating a requirement for both Nck and N-WASP, but not WIP or WIP family members in pathogen-induced actin assembly.
Curr Biol. 2012 Oct 23;22(20):1962-8
Gujas B, Alonso-Blanco C, Hardtke CS.
 Soil acidification is a major agricultural problem that negatively affects crop yield [ [1] and [2]]. Root systems counteract detrimental passive proton influx from acidic soil through increased proton pumping into the apoplast [3], which is presumably also required for cell elongation and stimulated by auxin [ [4] and [5]].
Soil acidification is a major agricultural problem that negatively affects crop yield [ [1] and [2]]. Root systems counteract detrimental passive proton influx from acidic soil through increased proton pumping into the apoplast [3], which is presumably also required for cell elongation and stimulated by auxin [ [4] and [5]].
Here, we found an unexpected impact of extracellular pH on auxin activity and cell proliferation rate in the root meristem of two Arabidopsis mutants with impaired auxin perception, axr3 and brx [ [6] and [7]]. Surprisingly, neutral to slightly alkaline media rescued their severely reduced root (meristem) growth by stimulating auxin signaling, independent of auxin uptake. The finding that proton pumps are hyperactive in brx roots could explain this phenomenon and is consistent with more robust growth and increased fitness of brx mutants on overly acidic media or soil. Interestingly, the original brx allele was isolated from a natural stock center accession collected from acidic soil [8].
Our discovery of a novel brx allele in accessions recently collected from another acidic sampling site demonstrates the existence of independently maintained brx loss-of-function alleles in nature and supports the notion that they are advantageous in acidic soil pH conditions, a finding that might be exploited for crop breeding.
Organizado por Manuel Fuentes, Concha Gil y el científico del CNB Juan Pablo Albar, los días 18 y 19 de octubre de 2012 se celebra en la Universidad Complutense de Madrid el III ProteoRed-ISCIII Protein Microarrays Course.
El curso se centra en un repaso a los conocimientos más actuales en el campo de Protein Microarrays, siendo una magnífica oportunidad para mejorar la competitividad de la unidades españolas de proteómica.
Tras el brote de E. coli que afectó a Alemania y posteriormente a Francia la primavera del año pasado, la Academia Europea de Microbiología reunió el pasado diciembre en París a un grupo de expertos a nivel mundial para analizar lo sucedido y proponer estrategias para evitar o al menos minimizar los efectos de otros posibles brotes. En la revista científica EMBO Molecular Medicine y coordinado por el investigador del Centro Nacional de Biotecnología del CSIC Miguel Vicente, se publica ahora un artículo que recoge las conclusiones de la reunión.
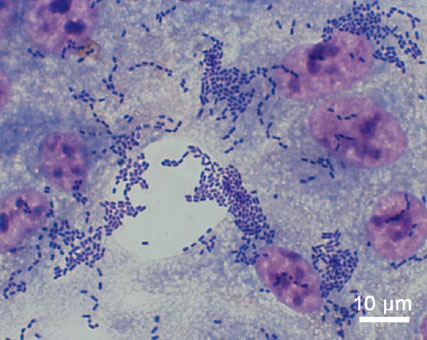 A pesar de la rapidez con la que se secuenció el genoma de la cepa de E. coli causante del brote, la mal llamada en España crisis de los pepinos se saldó con la muerte de 54 personas, en su mayoría ciudadanos alemanes, y una enormes pérdidas económicas en varios países, entre ellos España. Pese a todos los esfuerzos por identificarlo, no se tiene certeza de dónde ni cómo se originó el brote epidémico y solo existen explicaciones circunstanciales. Lamentablemente, un brote similar puede volver a darse en cualquier momento por lo que los científicos de la Academia Europea de Microbiología señalan una serie de recomendaciones para minimizar en lo posible sus efectos.
A pesar de la rapidez con la que se secuenció el genoma de la cepa de E. coli causante del brote, la mal llamada en España crisis de los pepinos se saldó con la muerte de 54 personas, en su mayoría ciudadanos alemanes, y una enormes pérdidas económicas en varios países, entre ellos España. Pese a todos los esfuerzos por identificarlo, no se tiene certeza de dónde ni cómo se originó el brote epidémico y solo existen explicaciones circunstanciales. Lamentablemente, un brote similar puede volver a darse en cualquier momento por lo que los científicos de la Academia Europea de Microbiología señalan una serie de recomendaciones para minimizar en lo posible sus efectos.
- La cepa O104:H4 muestra una sorprendente capacidad de adhesión al intestino y produce una toxina que provoca graves daños renales. Estos efectos por ahora sólo se pueden paliar eliminando la toxina mediante diálisis y manteniendo el equilibrio de electrolitos en sangre.
- Algunos antibióticos como meropenem, azitromicina o tigeciclina podrían utilizarse ya que la cepa es sensible a ellos. Otros antibióticos como la ciprofloxacina han de usarse con mucha precaución pues en dosis no inhibitorias pueden provocar un aumento en la producción de la toxina.
- Un anticuerpo comercializado como Eculizumab puede disminuir los efectos negativos de la respuesta inmunitaria que se genera en el organismo contra las toxinas de esta cepa de E. coli, pero tiene el riesgo asociado de disminuir la propia capacidad de defensa del cuerpo.
- Los expertos recomiendan que las autoridades sanitarias dispongan de procedimientos para que en este tipo de brotes el genotipado de las bacterias causantes se haga a la mayor brevedad posible. De el depende tanto el tratamiento como la eliminación de los focos de infección.
- Se hace asimismo hincapié en la necesidad de investigar a fondo los mecanismos por los que la toxina hace su efecto para así conseguir fármacos que impidan su acción en el riñón, y en la búsqueda de nuevos antibióticos que frenen la infección.
- Por último, proponen que se mejore la comunicación entre los científicos, los responsables políticos y los medios de difusión para que se afronten estos sucesos con una base científica fiable y adecuada a cada caso para minimizar los efectos sobre los ciudadanos y los sectores productivos.
EMBO Mol Med. 2012 Mar;4(3):160-4
García-del Portillo F, Cossart P.
 A workshop on ‘The Biology of Intracellular Bacterial Pathogens’ was held last October in a venue of the International University of Andalusia (UNIA) located in the World Historic Heritage town of Baeza, in the South of Spain. This Workshop gathered leading scientists from around the world to discuss their latest findings related to the mechanisms that intracellular pathogens use to subvert and manipulate host cell functions.
A workshop on ‘The Biology of Intracellular Bacterial Pathogens’ was held last October in a venue of the International University of Andalusia (UNIA) located in the World Historic Heritage town of Baeza, in the South of Spain. This Workshop gathered leading scientists from around the world to discuss their latest findings related to the mechanisms that intracellular pathogens use to subvert and manipulate host cell functions.
The workshop focused on novel aspects that imprint current research in this discipline, including the heterogeneous behaviour of the pathogen at the population level, the host determinants that modulate susceptibility to the infection, the search for new drugs to combat these particular types of infections and also cutting edge technologies based on new imaging approaches and the use of microfluidics. Discussion on these topics provided new insights into the biology of these pathogens and enriched the field with new ideas for understanding why colonization of the intracellular niche of eukaryotic cells is a preferred strategy used by important human pathogens.
Perez-Berna AJ, Ortega-Esteban A, Menendez-Conejero R, Winkler DC, Menendez M, Steven AC, Flint SJ, de Pablo PJ, San Martín C.
 Adenovirus assembly concludes with proteolytic processing of several
capsid and core proteins. Immature virions containing precursor proteins
lack infectivity because they cannot properly uncoat, becoming trapped
in early endosomes. Structural studies have shown that precursors
increase the network of interactions maintaining virion integrity. Using
different biophysical techniques to analyze capsid disruption in vitro,
we show that immature virions are more stable than the mature ones
under a variety of stress conditions, and that maturation primes
adenovirus for highly cooperative DNA release. Cryo-electron tomography
reveals that under mildly acidic conditions mimicking the early
endosome, mature virions release pentons and peripheral core contents.
At higher stress levels, both mature and immature capsids crack open.
Adenovirus assembly concludes with proteolytic processing of several
capsid and core proteins. Immature virions containing precursor proteins
lack infectivity because they cannot properly uncoat, becoming trapped
in early endosomes. Structural studies have shown that precursors
increase the network of interactions maintaining virion integrity. Using
different biophysical techniques to analyze capsid disruption in vitro,
we show that immature virions are more stable than the mature ones
under a variety of stress conditions, and that maturation primes
adenovirus for highly cooperative DNA release. Cryo-electron tomography
reveals that under mildly acidic conditions mimicking the early
endosome, mature virions release pentons and peripheral core contents.
At higher stress levels, both mature and immature capsids crack open.
The viral core is completely released from cracked capsids in mature
virions, but remains connected to shell fragments in the immature
particle. The extra stability of immature adenovirus does not equate
with greater rigidity, since in nanoindentation assays immature virions
exhibit greater elasticity than the mature particles. Our results have
implications for the role of proteolytic maturation in adenovirus
assembly and uncoating. Precursor proteins favor assembly by
establishing stable interactions with the appropriate curvature, and
preventing premature ejection of contents by tightly sealing the capsid
vertices. Upon maturation, core organization is looser, particularly at
the periphery, and interactions preserving capsid curvature are
weakened. The capsid becomes brittle, and pentons are more easily
released. Based on these results, we hypothesize that changes in core
compaction during maturation may increase capsid internal pressure to
trigger proper uncoating of adenovirus..
 Adenovirus (AdV) capsid organization is considerably complex, not only
because of its large size (~950 Å) and triangulation number (pseudo T =
25), but also because it contains four types of minor proteins in
specialized locations modulating the quasi-equivalent icosahedral
interactions. Up until 2009, only its major components (hexon, penton,
and fiber) had separately been described in atomic detail. Their
relationships within the virion, and the location of minor coat
proteins, were inferred from combining the known crystal structures with
increasingly more detailed cryo-electron microscopy (cryoEM) maps.
Adenovirus (AdV) capsid organization is considerably complex, not only
because of its large size (~950 Å) and triangulation number (pseudo T =
25), but also because it contains four types of minor proteins in
specialized locations modulating the quasi-equivalent icosahedral
interactions. Up until 2009, only its major components (hexon, penton,
and fiber) had separately been described in atomic detail. Their
relationships within the virion, and the location of minor coat
proteins, were inferred from combining the known crystal structures with
increasingly more detailed cryo-electron microscopy (cryoEM) maps.
There was no structural information on assembly intermediates. Later on
that year, two reports described the structural differences between the
mature and immature adenoviral particle, starting to shed light on the
different stages of viral assembly, and giving further insights into the
roles of core and minor coat proteins during morphogenesis.
Finally, in 2010, two papers describing the atomic resolution structure
of the complete virion appeared. These reports represent a
veritable tour de force for two structural biology techniques: X-ray
crystallography and cryoEM, as this is the largest macromolecular
complex solved at high resolution by either of them. In particular, the
cryoEM analysis provided an unprecedented clear picture of the complex
protein networks shaping the icosahedral shell. Here I review these
latest developments in the field of AdV structural studies.
Para poder seguir desarrollando la idea de sustituir a los antibióticos por virus que maten a las bacterias, en el Centro Nacional de Biotecnología del CSIC (CNB) estudian las proteínas que permiten a esos virus bacteriófagos anclarse sobre la superficie de las bacterias. Y acaban de publicar en la revista PNAS un avance realmente interesante, desvelando la estructura de una de estas proteínas.
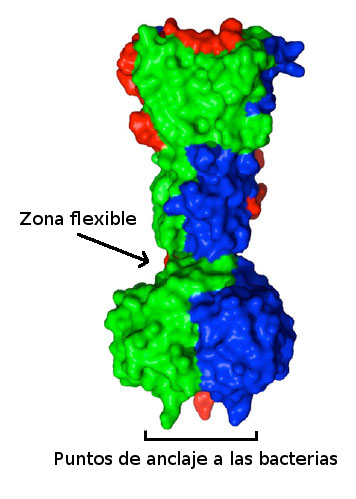 Cada bacteriófago se adhiere de forma muy específica a una especie concreta de bacteria, por lo que, en principio estos virus nunca podrían usarse de forma generalizada. Por ello, y como parte de un proyecto financiado por la Fundación Bill & Melinda Gates, en el grupo de Mark van Raaij trabajan sobre la idea de crear mediante mutaciones puntuales una gran variedad de bacteriófagos que puedan ser utilizados contra el tipo de bacteria que se desee. Su primer paso ha sido estudiar el mecanismo exacto por el que los bacteriófagos se colocan sobre la bacteria y se anclan a su membrana justo antes de empezar la destrucción de la misma.
Cada bacteriófago se adhiere de forma muy específica a una especie concreta de bacteria, por lo que, en principio estos virus nunca podrían usarse de forma generalizada. Por ello, y como parte de un proyecto financiado por la Fundación Bill & Melinda Gates, en el grupo de Mark van Raaij trabajan sobre la idea de crear mediante mutaciones puntuales una gran variedad de bacteriófagos que puedan ser utilizados contra el tipo de bacteria que se desee. Su primer paso ha sido estudiar el mecanismo exacto por el que los bacteriófagos se colocan sobre la bacteria y se anclan a su membrana justo antes de empezar la destrucción de la misma.
Analizando con precisión la estructura de las fibras mediante las que el bacteriófago T7 se une a las bacterias, Carmela García Doval y Mark van Raaij han comprobado que están formadas por tres unidades de una misma proteína. Mediante cristalografía de rayos X han determinado la existencia de un área justo antes de la zona que se une a las bacterias que dota a estas fibras de flexibilidad. Una flexibilidad que parece ser importante a la hora de que el virus se ancle correctamente sobre la pared bacteriana.
Gracias a la alta resolución de los datos obtenidos, han podido localizar con precisión las zonas concretas a través de las cuales se produce la unión entre el fago y la bacteria. Estos datos corroboran lo que venían indicando estudios previos de los aminoácidos que forman estas proteínas. Ahora, en su laboratorio del CNB, el grupo de van Raaij pretende generar bacteriófagos que contengan mutaciones aleatorias en las zonas que determinan su unión a las bacterias.
Con los miles de mutantes que planean obtener, tendrán que ir analizando la especificidad con la que se unen a las diferentes bacterias. Una vez que hayan detectado los mutantes que eliminan específicamente a las bacterias patógenas que les interesen, se producirán en grandes cantidades para ensayar su uso como posible tratamiento.
García-Doval C, van Raaij MJ. Structure of the receptor-binding carboxy-terminal domain of bacteriophage T7 tail fibers. Proc Natl Acad Sci USA. 2012 May 28.
Manipulando moléculas individuales de la ADN polimerasa del virus Phi29, el equipo de Borja Ibarra ha conseguido cuantificar por primera vez el mecanismo de apertura del ADN utilizado por esta proteína. Publicado en la revista PNAS, este trabajo ayudará a desarrollar nanomotores sintéticos.
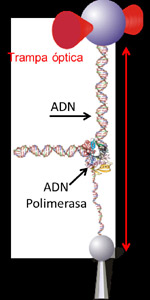 Muchas de las proteínas del interior celular funcionan como auténticos motores moleculares capaces de trabajar eficientemente a escalas nanométricas (mil veces más pequeñas que la de la célula). Para ello, transforman la energía disponible en el interior celular en diminutas fuerzas (picoNewtons: la diez millonésima parte del peso de un miligramo) y desplazamientos (nanómetros: la millonésima parte de un milímetro). Unos de los motores moleculares mas sorprendentes son las ADN polimerasas, las proteínas encargadas de duplicar la doble hélice del ADN que son capaces de leer la composición de bases de cada una de las hebras del ADN e incorporar la base complementaria en cada posición. Sorprendentemente, la polimerasa del virus Phi29 puede avanzar a lo largo del ADN a una velocidad de 6.000 bases por minuto a la vez que abre la doble hélice de la molécula y avanza por ella. Un corredor de obstáculos con estas propiedades sería capaz de correr aproximadamente a 360 km/h e ir saltando vallas separadas por solo 1 metro ¡sin tirar ninguna! Estas sorprendentes propiedades y habilidades de estos nanomotores biológicos, han fascinado durante mucho tiempo a los biólogos moleculares y en los últimos años a una amplia comunidad de físicos e investigadores del área de la nanotecnología.
Muchas de las proteínas del interior celular funcionan como auténticos motores moleculares capaces de trabajar eficientemente a escalas nanométricas (mil veces más pequeñas que la de la célula). Para ello, transforman la energía disponible en el interior celular en diminutas fuerzas (picoNewtons: la diez millonésima parte del peso de un miligramo) y desplazamientos (nanómetros: la millonésima parte de un milímetro). Unos de los motores moleculares mas sorprendentes son las ADN polimerasas, las proteínas encargadas de duplicar la doble hélice del ADN que son capaces de leer la composición de bases de cada una de las hebras del ADN e incorporar la base complementaria en cada posición. Sorprendentemente, la polimerasa del virus Phi29 puede avanzar a lo largo del ADN a una velocidad de 6.000 bases por minuto a la vez que abre la doble hélice de la molécula y avanza por ella. Un corredor de obstáculos con estas propiedades sería capaz de correr aproximadamente a 360 km/h e ir saltando vallas separadas por solo 1 metro ¡sin tirar ninguna! Estas sorprendentes propiedades y habilidades de estos nanomotores biológicos, han fascinado durante mucho tiempo a los biólogos moleculares y en los últimos años a una amplia comunidad de físicos e investigadores del área de la nanotecnología.
Clásicamente, el funcionamiento de estos motores se ha estudiado en tubos de ensayo donde millones de polimerasas trabajan al mismo tiempo en una reacción no sincronizada. De esta manera, muchos de los detalles del funcionamiento intrínseco de cada polimerasa se pierden en el promediado final. En colaboración con los grupo de José María Valpuesta y José L. Carrascosa en el Centro Nacional de Biotecnología del CSIC, de Francisco J. Cao en la Universidad Complutense de Madrid y de Margarita Salas del Centro de Biología Molecular “Severo Ochoa”, Borja Ibarra ha utilizado en su laboratorio del Instituto Madrileño de Estudios Avanzados en Nanocienca (IMDEA Nanociencia) la técnica de las pinzas ópticas para atrapar y manipular moléculas individuales de la polimerasa del virus Phi29. De esta forma estos investigadores han podido seguir la actividad de una sola molécula de polimerasa mientras trabaja y se desplaza abriendo la doble hélice del ADN.
En su estudio han encontrando que este diminuto motor molecular es capaz de acoplar la energía térmica con la energía química derivada de la incorporación de nucleótidos en la ruptura de los puentes de hidrógeno que mantienen unidas las dos cadenas del ADN. De este modo, se llegan a ejercer sobre la doble cadena de ADN fuerzas superiores a los 10 pN (1010 veces su peso). Escalando la polimerasa al tamaño microscópico, un motor del tamaño del de un coche con la misma relación fuerza-masa sería capaz de ejercer fuerzas efectivas iguales al peso de 300 millones de toneladas métricas, o el peso de unos 700 portaaviones.
Hace tan solo unos años la posibilidad de estudiar los sistemas biológicos a nivel de moléculas individuales parecía una historia sacada de algún libro de ciencia ficción, pero la implementación de avanzadas técnicas de manipulación de moléculas individuales tanto en el IMDEA Nanociencia como en el Centro Nacional de Biotecnología del CSIC está permitiendo que estas historias se hagan realidad. El estudio de los motores moleculares biológicos con estas técnicas esta permitiendo por primera vez en España el establecimiento de una amplia red de colaboración interdisciplinar entre biofísicos, biólogos moleculares y físicos. Además estos nuevos descubrimientos permiten conocer mejor el funcionamiento interno de las células y harán posible diseñar en el futuro nanomáquinas sintéticas que emulen la ingeniosa y eficiente maquinaria molecular diseñada por la naturaleza.
Morin JA, Cao FJ, Lázaro JM, Arias-Gonzalez JR, Valpuesta JM, Carrascosa JL, Salas M, Ibarra B. Active DNA unwinding dynamics during processive DNA replication. Proc Natl Acad Sci USA. 2012 May 9.
En colaboración con IMDEA Nanociencia, el investigador del CNB Dr. Fernando Moreno ha organizado el 2º Workshop en Nanobiociencia. Tendrá lugar el próximo viernes 18 de Mayo en la sede del IMDEA Nanociencia, en el campus de la Universidad Autónoma de Madrid.
 Esta jornada científica, de un día de duración y con entrada libre, presenta una pequeña muestra de la biociencia de molécula individual que se realiza en España usando técnicas biofísicas como Microscopía de Fuerzas Atómicas (AFM), Pinzas Ópticas y Magnéticas, o Microscopía de Fluorescencia de Alta resolución.
Esta jornada científica, de un día de duración y con entrada libre, presenta una pequeña muestra de la biociencia de molécula individual que se realiza en España usando técnicas biofísicas como Microscopía de Fuerzas Atómicas (AFM), Pinzas Ópticas y Magnéticas, o Microscopía de Fluorescencia de Alta resolución.
El programa incluye cuatro sesiones con dos conferencias plenarias de investigadores extranjeros expertos en AFM y Pinzas Ópticas y contribuciones cortas de jóvenes biofísicos españoles.
COOKIES POLICY
A cookie is a text file that is stored on your computer or mobile device via a web server and only that server will be able to retrieve or read the contents of the cookie and allow the Web site remember browser preferences and navigate efficiently. Cookies make the interaction between the user and the website faster and easier.
General information
This Website uses cookies. Cookies are small text files generated by the web pages you visit, which contain the session data that can be useful later in the website. In this way this Web remembers information about your visit, which can facilitate your next visit and make the website more useful.
How do cookies?
Cookies can only store text, usually always anonymous and encrypted. No personal information is ever stored in a cookie, or can be associated with identified or identifiable person.
The data allow this website to keep your information between the pages, and also to discuss how to interact with the website. Cookies are safe because they can only store information that is put there by the browser, which is information the user entered in the browser or included in the page request. You can not run the code and can not be used to access your computer. If a website encrypts cookie data, only the website can read the information.
What types of cookies used?
The cookies used by this website can be distinguished by the following criteria:
1. Types of cookies as the entity that manages:
Depending on who the entity operating the computer or domain where cookies are sent and treat the data obtained, we can distinguish:
- Own cookies: are those that are sent to the user's terminal equipment from a computer or domain managed by the editor itself and from which provides the service requested by the user.
- Third party cookies: these are those that are sent to the user's terminal equipment from a machine or domain that is not managed by the publisher, but by another entity data is obtained through cookies.
In the event that the cookies are installed from a computer or domain managed by the editor itself but the information collected by these is managed by a third party can not be considered as party cookies.
2. Types of cookies as the length of time that remain active:
Depending on the length of time that remain active in the terminal equipment can be distinguished:
- Session cookies: cookies are a type designed to collect and store data while the user accesses a web page. Are usually used to store information that only worth preserving for the service requested by the user at any one time (eg a list of products purchased).
- Persistent cookies: cookies are a type of data which are stored in the terminal and can be accessed and treated for a period defined by the head of the cookie, and can range from a few minutes to several years.
3. Cookies types according to their purpose:
Depending on the purpose for which the data are processed through cookies, we can distinguish between:
- Technical cookies: these are those that allow the user to navigate through a web page or application platform and the use of different options or services it exist as, for example, control traffic and data communication, identify the session, access to restricted access parts, remember the elements of an order, make the buying process an order, make an application for registration or participation in an event, use security features while browsing store content for dissemination videos or sound or share content via social networks.
- Customization cookies: these are those that allow the user to access the service with some general characteristics based on a predefined set of criteria in the user terminal would eg language, the type of browser through which you access the service, the locale from which you access the service, etc.
- Analysis cookies: they are those that allow the responsible for them, monitoring and analyzing the behavior of users of the web sites that are linked. The information gathered through such cookies are used in measuring the activity of web sites, application or platform and for the profiling of user navigation of such sites, applications and platforms, in order to make improvements function data analysis how users use the service.
Management tool cookies
This Website uses Google Analytics.
Google Analytics is a free tool from Google that primarily allows website owners know how users interact with your website. Also, enable cookies in the domain of the site in which you are and uses a set of cookies called "__utma" and "__utmz" to collect information anonymously and reporting of website trends without identifying individual users..
For statistics of use of this website use cookies in order to know the level of recurrence of our visitors and more interesting content. This way we can concentrate our efforts on improving the most visited areas and make the user more easily find what they are looking for. On this site you can use the information from your visit for statistical evaluations and calculations anonymous data and to ensure the continuity of service or to make improvements to their websites. For more details, see the link below privacy policy [http://www.google.com/intl/en/policies/privacy/]
How to manage cookies on your computer: disabling and deleting cookies
All Internet browsers allow you to limit the behavior of a cookie or disable cookies within settings or browser settings. The steps for doing so are different for each browser, you can find instructions in the help menu of your browser.
If you decline the use of cookies, since it is possible thanks to the preferences menu of your browser or settings, reject, this website will continue to function properly without the use of the same.
Can you allow, block or delete cookies installed on your computer by setting your browser options installed on your computer:
- For more information about Internet Explorer click here.
- For more information on Chrome click here.
- For more information about Safari click here.
- For more information about Firefox click here.
Through your browser, you can also view the cookies that are on your computer, and delete them as you see fit. Cookies are text files, you can open and read the contents. The data within them is almost always encrypted with a numeric key corresponding to an Internet session so often has no meaning beyond the website who wrote it.
Informed consent
The use of this website on the other hand, implies that you paid your specific consent to the use of cookies, on the terms and conditions provided in this Cookies Policy, without prejudice to the measures of deactivation and removal of cookies that you can take, and mentioned in the previous section.






