Vicente Rubio
Group Leader
Research summary
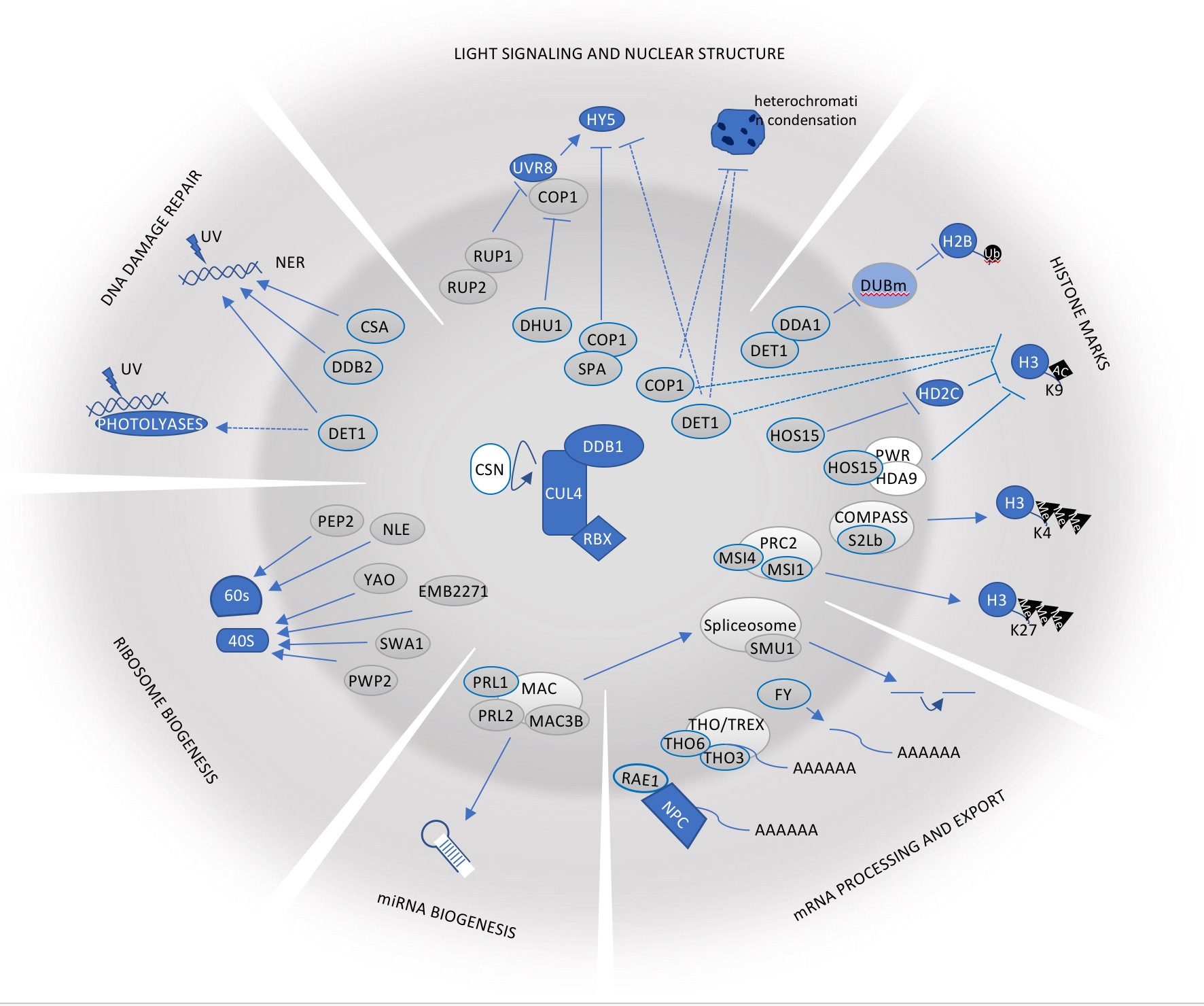
The group is focused on the characterization of new functions of the ubiquitination machinery in plants. For this, we study E3 ubiquitin ligases, such as CRL4-CDDD complexes, that act as hubs to integrate signals from different abiotic stresses and environmental stimuli in plants to trigger coherent adaptive and developmental responses. Interestingly, CRL4-CDDD complexes act in close proximity to the chromatin, controlling the stability and function of a plethora of chromatin remodeling machineries and transcription regulators, and therefore regulate transcriptional reprogramming of plants in response to environmental cues.
Publications
Blanco-Touriñán N, Legris M, Minguet EG, Costigliolo-Rojas C, Nohales MA, Iniesto E, García-Leόn M, Pacín M, Heucken N, Blomeier T, Locascio A, Černý M, Esteve-Bruna D, Díez-Díaz M, Brzobohatý B, Frerigmann H, Zurbriggen MD, Kay SA, Rubio V, Blázquez MA, Casal JJ, Alabadí D. COP1 destabilizes DELLA proteins in Arabidopsis. Proc Natl Acad Sci U S A. 2020 Jun 16;117(24):13792-13799.
Chico JM, Lechner E, Fernandez-Barbero G, Canibano E, García-Casado G, Franco-Zorrilla JM, Hammann P, Zamarreño AM, García-Mina JM, Rubio V, Genschik P, Solano R. CUL3BPM E3 ubiquitin ligases regulate MYC2, MYC3, and MYC4 stability and JA responses. Proc Natl Acad Sci U S A. 2020 Mar 2. pii: 201912199.
Fonseca S, Rubio V. Arabidopsis CRL4 Complexes: Surveying Chromatin States and Gene Expression. Front Plant Sci. ,2019 Sep 17;10:1095.
García-León M, Cuyas L, El-Moneim DA, Rodriguez L, Belda-Palazón B, Sanchez-Quant E, Fernández Y, Roux B, Zamarreño ÁM, García-Mina JM, Nussaume L, Rodriguez PL, Paz-Ares J, Leonhardt N, Rubio V. Arabidopsis ALIX Regulates Stomatal Aperture and Turnover of Abscisic Acid Receptors. Plant Cell. 2019 Oct;31(10):2411-2429.
Nassrallah A, Rougée M, Bourbousse C, Drevensek S, Fonseca S, Iniesto E, Ait-Mohamed O, Deton-Cabanillas AF, Zabulon G, Ahmed I, Stroebel D, Masson V, Lombard B, Eeckhout D, Gevaert K, Loew D, Genovesio A, Breyton C, De Jaeger G, Bowler C, Rubio V, Barneche F. DET1-mediated degradation of a SAGA-like deubiquitination module controls H2Bub homeostasis. Elife. 2018 Sep 7;7. pii: e37892
 The relevance of protein ubiquitination as an integral mechanism of many signaling pathways in plants has been demonstrated extensively. Ubiquitin (Ub) conjugation to proteins (i.e. ubiquitination) may trigger degradation of protein targets at the 26S proteasome or changes in their properties (e.g., protein activity, localization, assembly and interaction ability), depending on the extent or specific Ub chain configurations. Protein ubiquitination is mediated by an enzymatic cascade in which different types of E3 Ub ligases provide the substrate specificity.
The relevance of protein ubiquitination as an integral mechanism of many signaling pathways in plants has been demonstrated extensively. Ubiquitin (Ub) conjugation to proteins (i.e. ubiquitination) may trigger degradation of protein targets at the 26S proteasome or changes in their properties (e.g., protein activity, localization, assembly and interaction ability), depending on the extent or specific Ub chain configurations. Protein ubiquitination is mediated by an enzymatic cascade in which different types of E3 Ub ligases provide the substrate specificity.
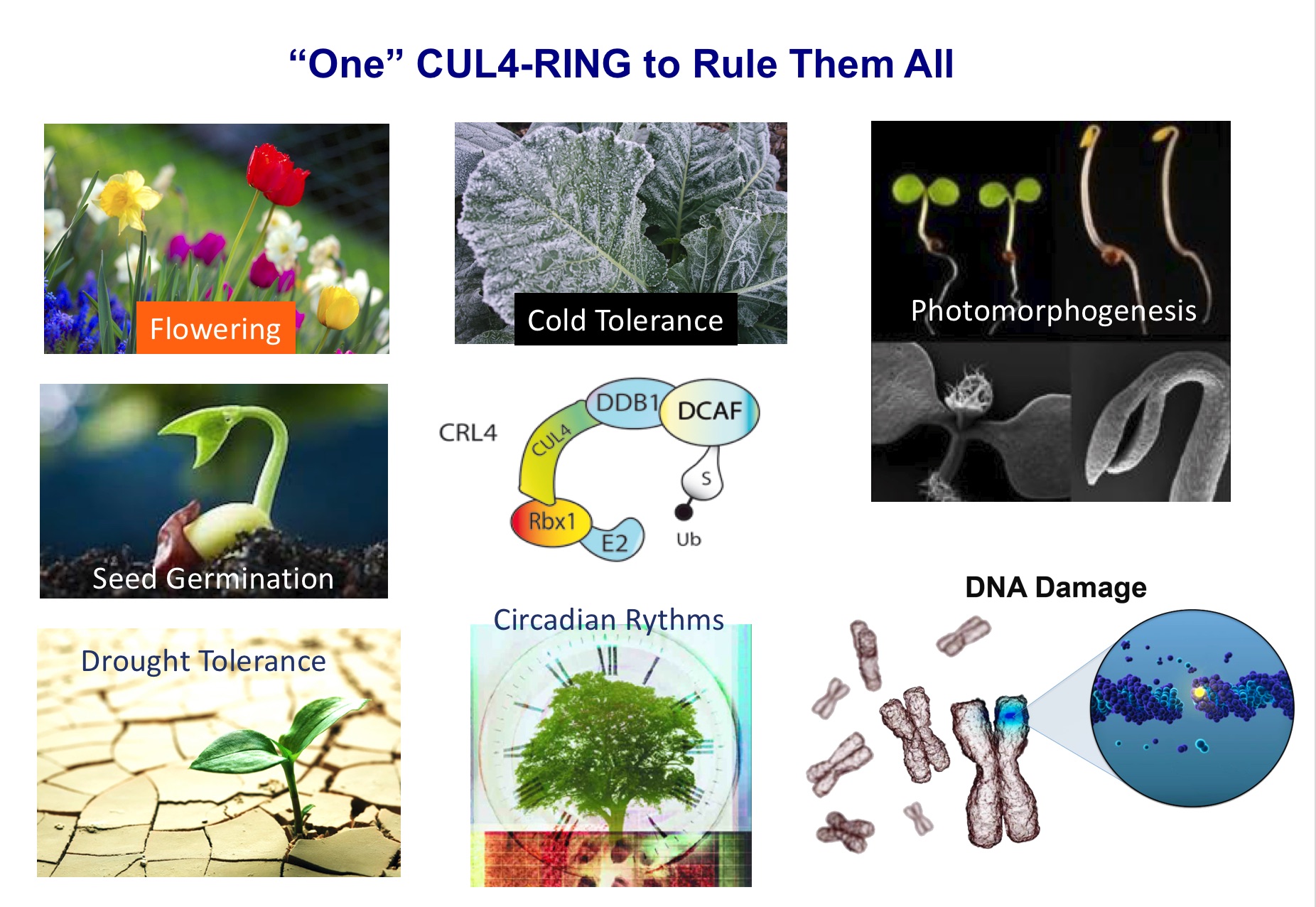
Among them, Cullin4 RING E3 ubiquitin ligases (CRL4) have been involved in biological processes spanning the plant´s whole life, including embryogenesis, seedling photomorphogenesis, circadian clock function, flowering and tolerance to different stresses (i.e. drought, high salinity, cold, osmotic stress) by promoting degradation of specific targets controlling those processes (Fig. 1). As an example, we have recently shown that DDA1, a substrate adaptor of CRL4-CDDD complexes, recognizes abscisic acid (ABA) receptors, triggering their ubiquitination and proteasomal degradation (Irigoyen et al., The Plant Cell 2014). Therefore, CRL4-CDDD complexes act as repressors of ABA-mediated water stress responses under optimal growth conditions. Interestingly, CRL4-CDDD function is performed in close proximity to chromatin, which should enable rapid translation of environmental and stress signals into changes in gene expression (Fig. 2; Fonseca and Rubio, 2019). Indeed, recent results from our laboratory showed that CRL4-CDDD complexes are part of a molecular pathway controlling epigenetic homeostasis (including Histone2B ubiquitination) in response to external stimuli (i.e. light conditions; Nassrallah et al., eLife 2018). Our current objectives aim to identify and characterize additional mechanisms by which CRL4-CDDD controls the accumulation of specific epigenetic marks over the plant genome in response to environmental changes, to regulate expression of specific set of genes that lead to plant adaptation to changing climate conditions.