Roberto Solano
Group Leader
Research summary
We are interested in understanding how plants are able to perceive changes in their environmentand integrate stress signals with their internal developmental programs, to induce adaptive responses and survive in nature. This integration depends on complex signalling networks that regulate the genetic reprogramming of the cell. The main focus of my lab is to understand one of the pathways involved in this network, the jasmonate(JA) signalling pathway, in plants.
Publications
Peñuelas M, Monte I, Schweizer F, Vallat A, Reymond P, García-Casado G, Franco-Zorrilla JM, Solano R. Jasmonate-Related MYC Transcription Factors Are Functionally Conserved in Marchantia polymorpha. The Plant Cell 2019; DOI: https://doi.org/10.1105/tpc.18.00974
Monte I, Franco-Zorrilla JM, García-Casado G, Angel M. Zamarreño, José M. García-Mina, Ryuichi Nishihama, Takayuki Kohchi and Roberto Solano. A Single JAZ Repressor Controls the Jasmonate Pathway in Marchantia polymorpha. Mol Plant. 2019;12(2):185‐198. doi:10.1016/j.molp.2018.12.017
Gimenez-Ibanez S, Zamarreño AM, García-Mina JM, Solano R. An Evolutionarily Ancient Immune System Governs the Interactions between Pseudomonas syringae and an Early-Diverging Land Plant Lineage. Current Biology 2019;29(14):2270‐2281.e4. doi:10.1016/j.cub.2019.05.079
Ortigosa A, Gimenez-Ibañez S, Leonhardt N and Solano R. Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnology Journal . 2019. 17, 665-673
Chico JM, Lechner E, Fernandez-Barbero G, Esther Canibano, Gloria García-Casado, Jose Manuel Franco-Zorrilla JM, Hammann P, Zamarreño AM, García-Mina JM, Rubio V, Genschik P, Solano R CUL3BPM E3 ubiquitin ligases regulate MYC2, MYC3, and MYC4 stability and JA responses. Proc Natl Acad Sci USA 2020;117(11):6205‐6215. doi:10.1073/pnas.1912199117
Monte I, Kneeshaw S, Franco-Zorrilla JM, Chini A, Zamarreño AM, García-Mina JM, Solano R. An Ancient COI1-Independent Function for Reactive Electrophilic Oxylipins in Thermotolerance. Curr Biol 2020; 30, 962–971 doi.org/10.1016/j.cub.2020.01.023
Ortigosa A, Fonseca S, Franco‐Zorrilla JM, Fernández‐Calvo P, Zander M, Lewsey MG, García‐Casado G, Fernández‐Barbero G, Ecker JR, Solano R. The JA‐pathway MYC transcription factors regulate photomorphogenic responses by targeting HY5 gene expression. The Plant Journal 2020 DOI: 10.1111/tpj.14618
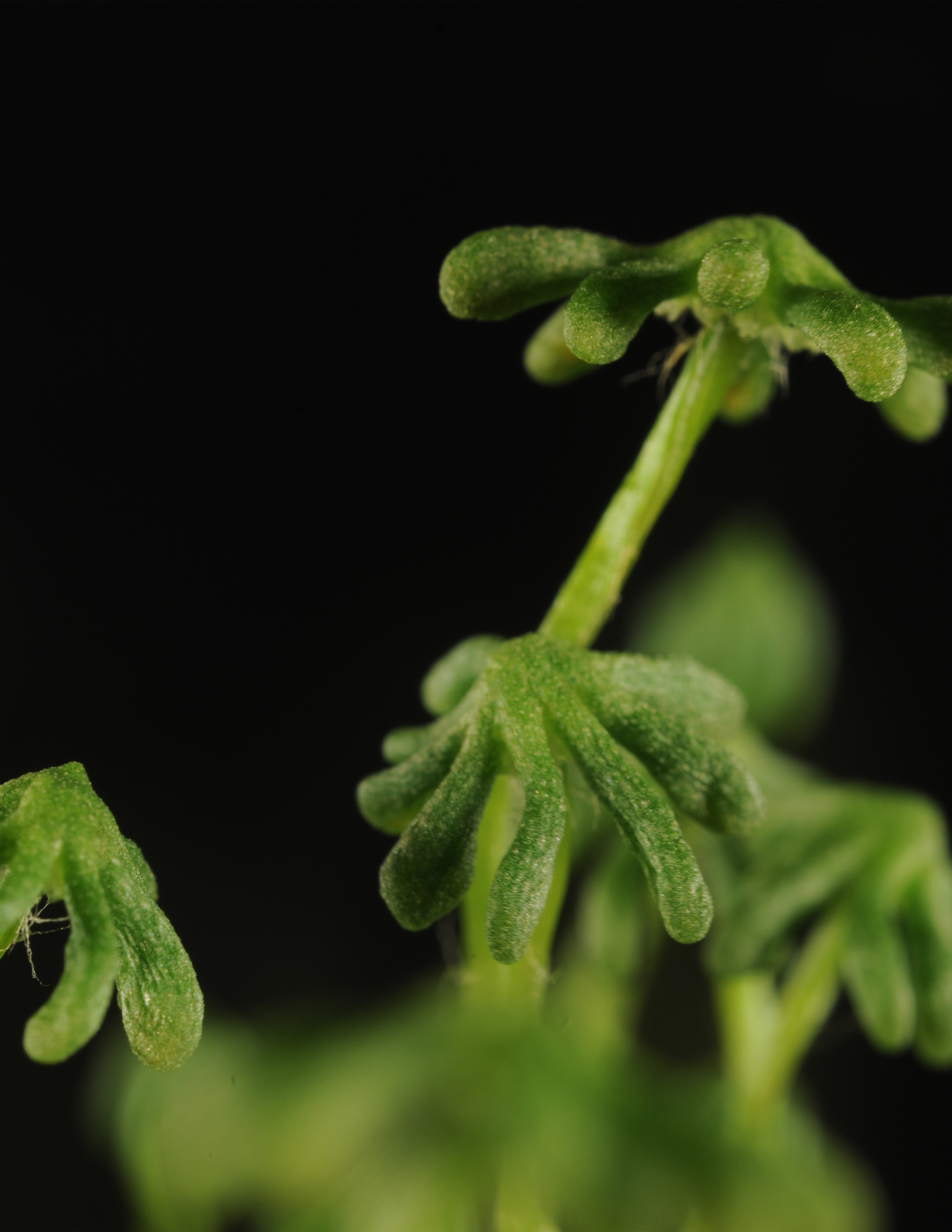

Jasmonates (JAs) are fatty acid-derived signalling molecules essential for the survival of plants in nature since they are important activators of stress responses and developmental programs. The main focus of my lab is to understand mechanistically the JA signalling pathway in plants; knowledge that is basic to design biotech and agronomical applications that improve plant resistance to stresses and plant yield. We have traditionally worked in the model plant Arabidopsis thaliana, but have recently focused in the Liverwort Marchantia polymorpha due to its remarkable genetic advantages, such as very low gene redundancy.
Our major achievements in the last two years are:
- Identification of a new pathway for thermotolerance in plants (Monte et al, Current Biology, 2020)
- Identification of CUL3BPM E3 ubiquitin ligases that regulate MYC transcription factors stability and JA responses (Chico et al, PNAS, 2020)
- Characterisation of conserved basal defence mechanisms in land plants (Gimenez-Ibanez et al, Current Biology, 2019)
- Design and obtention of a tomato resistant to bacterial speck by CRISPR/Cas9-based mutation of SlJAZ2 (Ortigosa et al, Plant Biotechnology Journal, 2019)
- Characterisation of MYC2 orthologs in Marchantia polymorpha (Peñuelas et al, The Plant Cell, 2019)
- Characterisation of the single JAZ repressor in Marchantia polymorpha (Monte et al, Molecular Plant, 2019)
- Identification of a new function of MYCs in photomorphogenesis (Ortigosa et al, The Plant Journal, 2020)
- Collaborated in the characterisation of PIF transcription factors in reproductive development (Costa Galvāo et al, Nature Communications, 2019)
- Discovery of bioactive hydroxylated derivatives of JA-Ile (Jimenez-Aleman et al, Biochim Biophys Acta Mol Cell Biol Lipids 2019)
- Identified the DNA target sequence of many plant transcription factors using previously developed tools and in collaboration with several groups (Ramírez Gonzales et al, The Plant Journal, 2020).
.