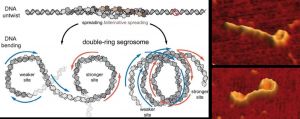
Certain bacteria maintain small DNA molecules that encode neurotoxic proteins. These DNA molecules utilize the cell machinery of the bacterial host to replicate and spread during the cell-division process. As explained in a study published in the journal Nucleic Acids Research, in order for these toxic DNA molecules to segregate, the bacteria fold the DNA into a double-ring shape, an architecture never observed until now.
Together with certain proteins, this double-ring DNA forms a structure called a segrosome and then becomes bound to a motor protein that moves genetic material to different points within the cell. To observe these structures, the investigators first used atomic force microscopy and then applied the technique called Magnetic Tweezers, which allowed them to study the formation of the double-ring structure in real time. The study was carried out using DNA from pathogen bacteria of the species Clostridium botulinum.
The work is a joint initiative of researchers from the Spanish National Center for Biotechnology (CNB) and the Biological Research Center (CIB), both part of the CSIC.
“The first step in the formation of this double ring takes place when the protein TubR links to a region of the DNA to form the segrosome,” explains Fernando Moreno, researcher at the CNB-CSIC and one of the leading researchers of the study. As a result, DNA bending takes place, giving way to the double-ring structures we observed using atomic force microscopy. “Lastly,” he continues, “the motor protein TubZ binds to the segrosome to move the genetic material to different points within the cell.”
According to Alejandro Martín-González, a fellow CNB-CSIC researcher and one of the study’s coauthors, “Doing research to understand how virulence plasmids segregate can lead us to discover new ways of preventing toxin production.”
.
- Martín-García B, Martín-González A, Carrasco C, Hernández-Arriaga AM, Ruíz-Quero R, Díaz-Orejas R, Aicart-Ramos C, Moreno-Herrero F, Oliva MA.. The TubR-centromere complex adopts a double-ring segrosome structure in Type III partition systems. Nucleic Acids Res. 2018 May 14. doi: 10.1093/nar/gky370