
José Ramón Valverde Carrillo
Group Leader
Service Description
 The Scientific Computing Service provides advanced support for scientific data analysis in bioinformatics and biocomputing through close collaboration and organisation of international courses (in-house and abroad).
The Scientific Computing Service provides advanced support for scientific data analysis in bioinformatics and biocomputing through close collaboration and organisation of international courses (in-house and abroad).
Our main areas of work include modern bioinformatics next-generation sequencing (NGS) data analyses, metagenomics and de novo genome sequencing for topics such as the effect of pesticides on rhizosphere, sequencing novel Escherichia coli mutant strains, as well as offering courses on phylogeny analysis and metagenomics.
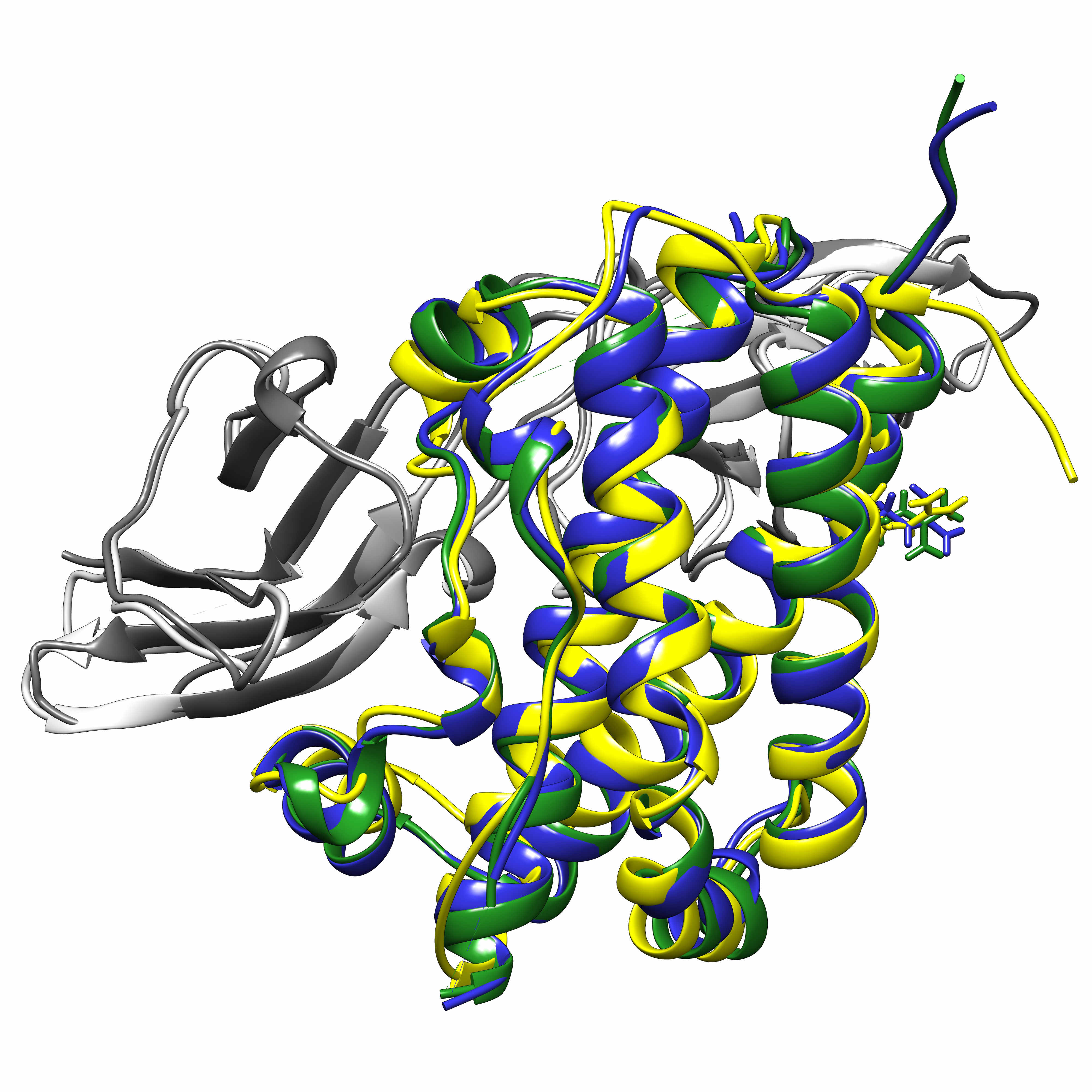
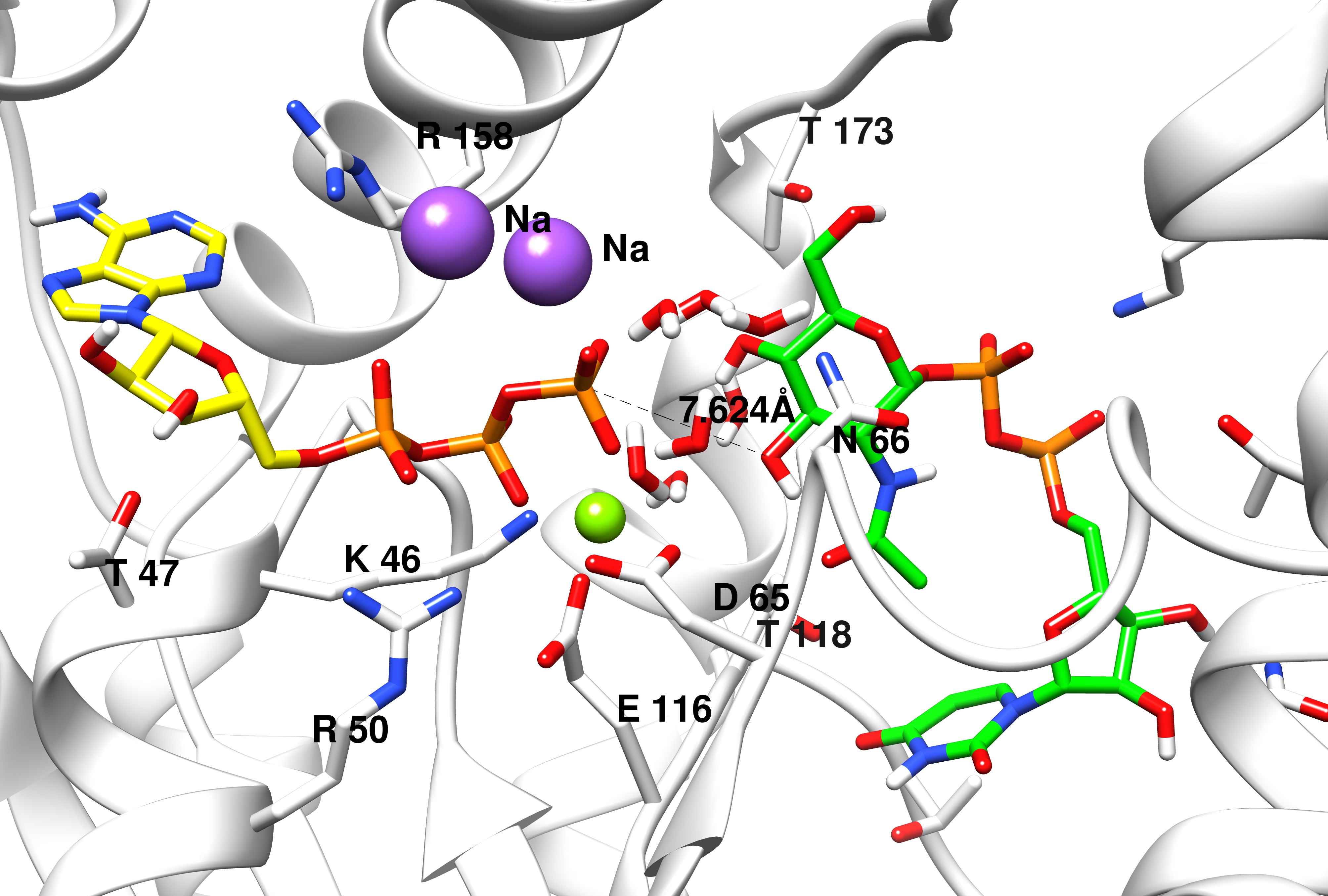
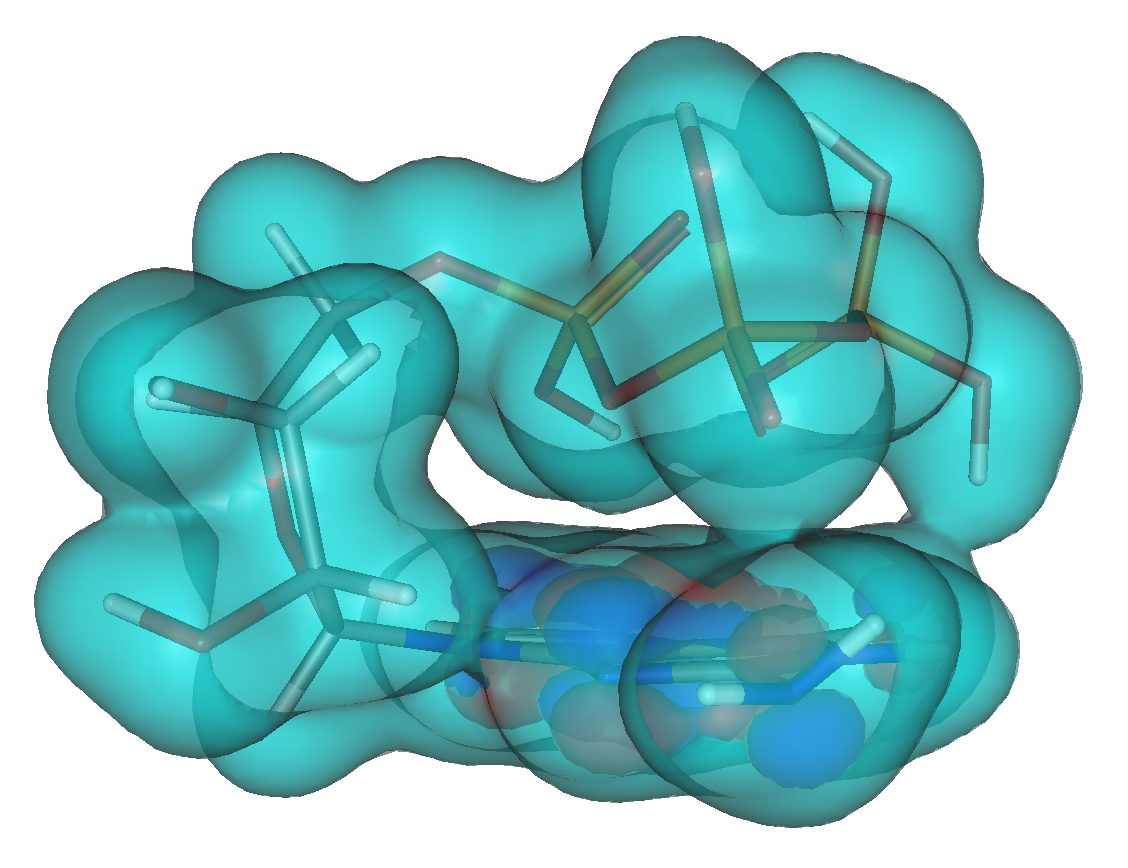
Computational biology includes binding pocket identification, docking, drug screening, in silico mutagenesis, molecular and quantum dynamics, QM and QM/MM (quantum mechanics/molecular mechanics) models and reaction modelling, which we use to study human growth hormone mutants, select natural compounds against cancer stem cells, resolve the reactions in the bacterial e-z complex; we also offer courses on biocomputing.
Advanced biostatistical analyses include treatment of complex experimental setups related to new therapies, proteomics, non-linear longitudinal analyses, and an international course on biostatistics.
We provide support for many computer languages to produce dedicated software and organise an international course on the Python programming language.
We coordinated the Iberoamerican Network on FLOSS for Biomedicine (CYTED 510RT0391), in collaboration with pioneering NGS groups, through COST action BM1006 (SEQAHEAD). We participate in EU CBRN CoE P35 and in EMBnet, and with numerous external institutions including the University of Alcalá de Henares, we coordinate the Master in Bioinformatics of the University of Colombo (Sri Lanka), the Advanced Course in Genomics organized by ILRI in Nairobi, Kenya, and many other courses in Europe, Latin America, Asia and Africa.
Fees and others





