In order to gain insight of the mechanisms behind these events, Rafael Giraldo's laboratory at the National Center for Biotechnology (CNB-CSIC), have developed a model to study the dynamics and transmission mechanisms of prions. Using synthetic biology techniques and the bacterial protein RepA–WH1, they had modified the conformational dynamics of the protein using various types of ligands such as DNA, phospholipids, or nanoparticles to induce its assembly into amyloid fibers. This model allows the study of protein toxicity in Escherichia coli, in which the original protein is harmless. The synthetic prion spreads in the form of two different structural variants during cell division. The most cytotoxic lineage produces pores in the bacterial inner membrane making it unable to detoxify free radicals that are generated as a result of membrane damage, resulting in bacterial death.
The journal mBio has just published their new work, where they have studied the transmission of the bacterial synthetic prion RepA-WH1 in cultured human and mice cell lines. Using several approaches, they have shown that prion propagation is only possible if the target cells previously contain the RepA-WH1 protein itself. In this case, the amyloid aggregates get inside the receptor cells, where they reproduce the structure on the soluble RepA-WH1 molecules, causing the formation of new intracellular amyloid aggregates. Giraldo highlights “the aggregated protein especially affects the functionality of mitochondria, the organelles responsible for cellular respiration, following an analogous mechanisms to those we had previously described in E. coli. This suggests that the mechanisms of transmission and amyloid cytotoxicity are universal and can be reconstructed by Synthetic Biology”. In addition, Giraldo remarks that one of the advantages of working with this synthetic prion is its biosecurity, since it does not spread in human or animal cells that do not naturally express RepA-WH1, allowing to understand their transmission dynamics without risks.
Prion transmission is an evolutionary conserved mechanism
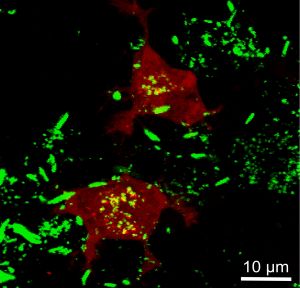 Prions (stained in green) promote the formation of intracellular amyloid aggregates in target cells
Aida Revilla, CNB-CSIC
Prions (stained in green) promote the formation of intracellular amyloid aggregates in target cells
Aida Revilla, CNB-CSIC
- The use of Synthetic Biology techniques allows a better understanding of prion toxicity and transmission mechanisms
- This new research study indicates that bacterial prions have the potential to be transmitted in human and mice cell lines
In most cases, cellular proteins need to acquire a specific three-dimensional (3D) structure to work properly. In mammals, there are severe pathologies that are associated with an incorrect 3D protein structure, termed amyloid, with changes that affect their functionality, and diseases caused by prions belong to this group. Prions are proteins that, in their normal conformation, have a non-damaging function for the cell. However, some switches in their folding structure give them the ability to transmit this misfolded shape onto normal variants of the same protein, aggregating into fibers that can also be transmitted between cells. Initially associated with Creutzfeldt-Jacob disease in humans or spongiform encephalopathy in cattle, now proteins involved in other neurodegenrative diseases, such as Alzheimer’s, Parkinson’s, and ELA, are also considered to be prions.
But this characteristic aggregation of prions does not occur only in mammals. In bacteria it has been shown that some proteins have this ability too, forming protein aggregates that are transmitted during cell division, and some studies suggest that, at least in vitro, they can be transmitted to other organisms.
COOKIES POLICY
A cookie is a text file that is stored on your computer or mobile device via a web server and only that server will be able to retrieve or read the contents of the cookie and allow the Web site remember browser preferences and navigate efficiently. Cookies make the interaction between the user and the website faster and easier.
General information
This Website uses cookies. Cookies are small text files generated by the web pages you visit, which contain the session data that can be useful later in the website. In this way this Web remembers information about your visit, which can facilitate your next visit and make the website more useful.
How do cookies?
Cookies can only store text, usually always anonymous and encrypted. No personal information is ever stored in a cookie, or can be associated with identified or identifiable person.
The data allow this website to keep your information between the pages, and also to discuss how to interact with the website. Cookies are safe because they can only store information that is put there by the browser, which is information the user entered in the browser or included in the page request. You can not run the code and can not be used to access your computer. If a website encrypts cookie data, only the website can read the information.
What types of cookies used?
The cookies used by this website can be distinguished by the following criteria:
1. Types of cookies as the entity that manages:
Depending on who the entity operating the computer or domain where cookies are sent and treat the data obtained, we can distinguish:
- Own cookies: are those that are sent to the user's terminal equipment from a computer or domain managed by the editor itself and from which provides the service requested by the user.
- Third party cookies: these are those that are sent to the user's terminal equipment from a machine or domain that is not managed by the publisher, but by another entity data is obtained through cookies.
In the event that the cookies are installed from a computer or domain managed by the editor itself but the information collected by these is managed by a third party can not be considered as party cookies.
2. Types of cookies as the length of time that remain active:
Depending on the length of time that remain active in the terminal equipment can be distinguished:
- Session cookies: cookies are a type designed to collect and store data while the user accesses a web page. Are usually used to store information that only worth preserving for the service requested by the user at any one time (eg a list of products purchased).
- Persistent cookies: cookies are a type of data which are stored in the terminal and can be accessed and treated for a period defined by the head of the cookie, and can range from a few minutes to several years.
3. Cookies types according to their purpose:
Depending on the purpose for which the data are processed through cookies, we can distinguish between:
- Technical cookies: these are those that allow the user to navigate through a web page or application platform and the use of different options or services it exist as, for example, control traffic and data communication, identify the session, access to restricted access parts, remember the elements of an order, make the buying process an order, make an application for registration or participation in an event, use security features while browsing store content for dissemination videos or sound or share content via social networks.
- Customization cookies: these are those that allow the user to access the service with some general characteristics based on a predefined set of criteria in the user terminal would eg language, the type of browser through which you access the service, the locale from which you access the service, etc.
- Analysis cookies: they are those that allow the responsible for them, monitoring and analyzing the behavior of users of the web sites that are linked. The information gathered through such cookies are used in measuring the activity of web sites, application or platform and for the profiling of user navigation of such sites, applications and platforms, in order to make improvements function data analysis how users use the service.
Management tool cookies
This Website uses Google Analytics.
Google Analytics is a free tool from Google that primarily allows website owners know how users interact with your website. Also, enable cookies in the domain of the site in which you are and uses a set of cookies called "__utma" and "__utmz" to collect information anonymously and reporting of website trends without identifying individual users..
For statistics of use of this website use cookies in order to know the level of recurrence of our visitors and more interesting content. This way we can concentrate our efforts on improving the most visited areas and make the user more easily find what they are looking for. On this site you can use the information from your visit for statistical evaluations and calculations anonymous data and to ensure the continuity of service or to make improvements to their websites. For more details, see the link below privacy policy [http://www.google.com/intl/en/policies/privacy/]
How to manage cookies on your computer: disabling and deleting cookies
All Internet browsers allow you to limit the behavior of a cookie or disable cookies within settings or browser settings. The steps for doing so are different for each browser, you can find instructions in the help menu of your browser.
If you decline the use of cookies, since it is possible thanks to the preferences menu of your browser or settings, reject, this website will continue to function properly without the use of the same.
Can you allow, block or delete cookies installed on your computer by setting your browser options installed on your computer:
- For more information about Internet Explorer click here.
- For more information on Chrome click here.
- For more information about Safari click here.
- For more information about Firefox click here.
Through your browser, you can also view the cookies that are on your computer, and delete them as you see fit. Cookies are text files, you can open and read the contents. The data within them is almost always encrypted with a numeric key corresponding to an Internet session so often has no meaning beyond the website who wrote it.
Informed consent
The use of this website on the other hand, implies that you paid your specific consent to the use of cookies, on the terms and conditions provided in this Cookies Policy, without prejudice to the measures of deactivation and removal of cookies that you can take, and mentioned in the previous section.





