
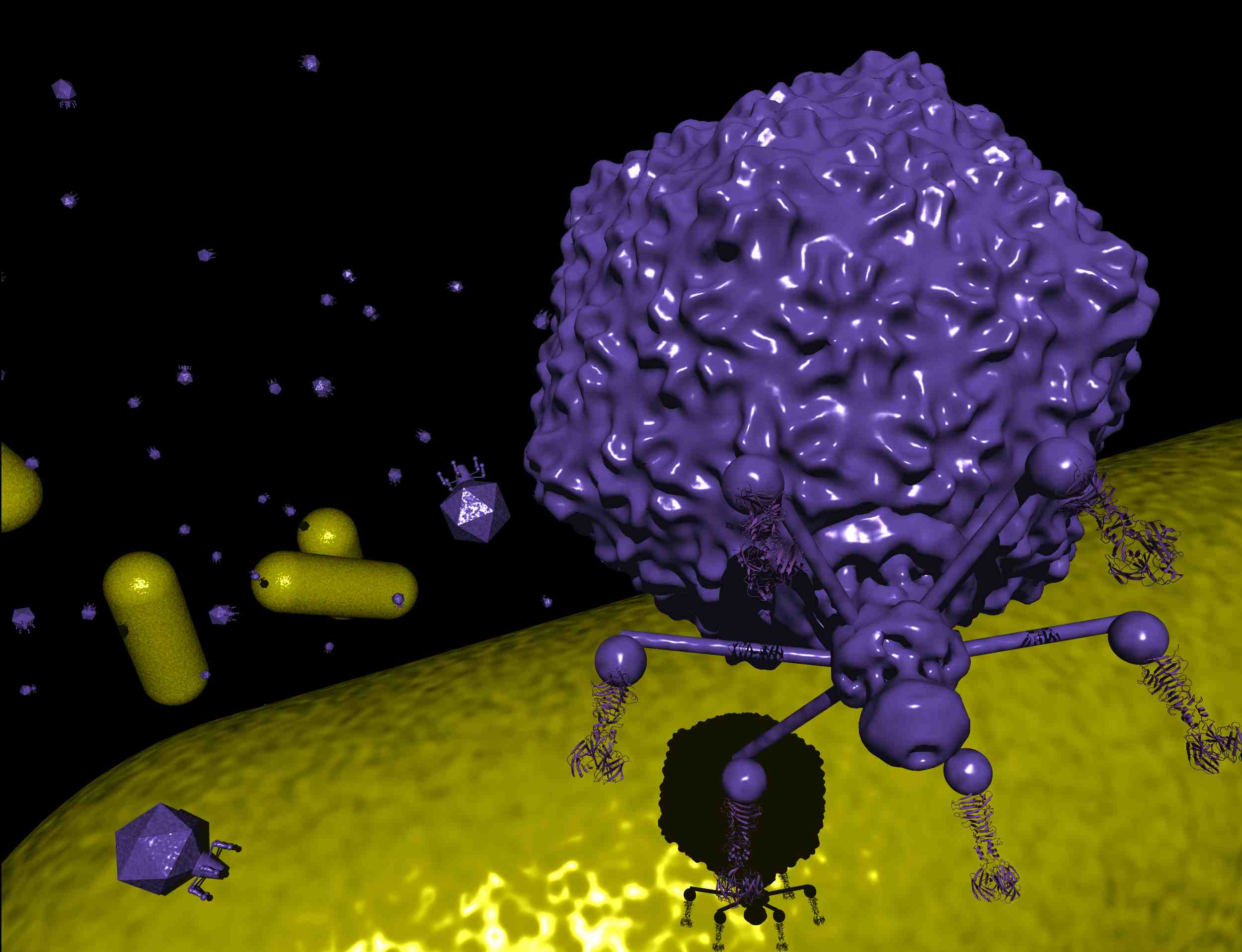
images courtesy of Florencio Pazos (left and right) and Marta Sanz (middle), CNB
structure
determination of bacteriophage proteins
supported by grant BFU2014-53425-P from the Spanish Ministery of Economy and Competiveness (MINECO)



images courtesy of Florencio Pazos (left and
right) and Marta Sanz (middle), CNB
Bacteriophages T4, T5 and T7 all
recognize host bacteria via fiber proteins. The T4 long tail
fibers are a complex of a trimer of gene product (gp) 34 (the
"thigh"), a monomer of gp35 (the "knee"), a trimer of gp36 and a
trimer of gp37, the last two together forming the "shin" and
receptor-binding "foot" or tip. We have recently determined the
structure of the tip of gp37 and have shown it contains a new
iron-binding fold. Recently, we have solved the structure of
a C-terminal fragment of gp34, which revealed a long triple
beta-helix.
We also solved the structure of a C-terminal fragment of the fiber
of phage T7, which revealed a modular fold composed of the pyramid
and tip domain.
The structure of the bacteriophage T5 L-shaped tail fiber tip,
with and without its chaperone, was also solved, giving us
examples of structure of fiber proteins of the Myoviridae,
Podoviridae and Siphoviridae families of the
Caudovirales order.
In collaboration with other groups, we study the physical
properties of the fiber proteins and their receptor-recognizing
ability.
publications
- Taylor NMI, van Raaij MJ, Leiman PG (2018)
Contractile injection systems of bacteriophages and related
systems. Mol Microbiol, in press.
- Hyman P,
van Raaij MJ (2018) Bacteriophage T4 long tail fiber
domains. Biophys Rev, in press.
- Granell M, Namura M, Alvira S, Kanamaru S, van Raaij MJ
(2017) Crystal structure of the carboxy-terminal region of
the bacteriophage T4 proximal long tail fiber protein gp34.
Viruses 9, 168.
- Garcia-Doval C, Caston JR,
Luque D, Granell M, Otero JM, Llamas-Saiz AL, Renouard
M, Boulanger P, van Raaij MJ (2015) Structure of the
receptor-binding carboxy-terminal domain of the
bacteriophage T5 L-shaped tail fibre with and without
Its intra-molecular chaperone. Viruses 7, 6424-6440.
- Gonzalez-Garcia VA, Pulido-Cid
M, Garcia-Doval C, Bocanegra R, van Raaij MJ, Martín-Benito J,
Cuervo A, Carrascosa JL (2015). Conformational changes leading to
T7 DNA delivery upon interaction with the bacterial receptor. J
Biol Chem. 290, 10038-10044.
- Granell
M, Namura M, Alvira S, Garcia-Doval C, Singh AK, Gutsche I, van
Raaij MJ, Kanamaru S (2014). Crystallization of the
carboxy-terminal region of the bacteriophage T4 proximal long tail
fibre protein gp34. Acta Cryst F70, 970-975.
- Ares P,
Garcia-Doval C, Llauro A, Gomez-Herrero J, van Raaij MJ, De
Pablo P (2014) The interplay between the mechanical
properties of bacteriophage fibers and the strength of
virus-host links during early infection stages. Phys Rev
E89, 052710.
- Kryshtafovych
A, Moult J, Bales P, Bazan JF, Biasini M, Burgin A, Chen C,
Cochran FV, Craig TK, Das R, Fass D, Garcia-Doval C, Herzberg O,
Lorimer D, Luecke H, Ma X, Nelson DC, van Raaij MJ, Rohwer F,
Segall A, Seguritan V, Zeth K, Schwede T (2014) Challenging the
state-of-the-art in protein structure prediction: Highlights of
experimental target structures for the 10th Critical Assessment of
Techniques for Protein Structure Prediction Experiment CASP10.
Proteins 82, Suppl 2, 26-42.
- Garcia-Doval C, Luque D,
Caston JR, Boulanger P, van Raaij MJ (2013) Crystallization of
the C-terminal domain of the bacteriophage T5 L-shape fibre.
Acta Cryst F69, 1363-1367.
- Cuervo A, Pulido-Cid
M, Chagoyen M, Arranz R, González-García VA, Garcia-Doval C,
Castón JR, Valpuesta JM, van Raaij MJ, Martín-Benito J,
Carrascosa JL (2013) Structural characterization of the
bacteriophage T7 tail machinery. J Biol Chem 288, 26290-26299.
- Garcia-Doval C, van Raaij MJ (2012) Structure of the
receptor-binding carboxy-terminal domain of bacteriophage T7
fibers. PNAS USA 109, 9390-9395.
- Garcia-Doval C, van Raaij MJ (2012) Crystallisation of
C-terminal domain of the bacteriophage T7 fibre gp17. Acta Cryst
F68, 166-171.
- Kryshtafovych A, Moult J, Bartual SG, Bazan JF, Berman H,
Casteel DE, Christodoulou E, Everett JK, Hausmann J, Heidebrecht
T, Hills T, Hui R, Hunt JF, Seetharaman J, Joachimiak A, Kennedy
MA, Kim C, Lingel A, Michalska K, Montelione GT, Otero JM,
Perrakis A, Pizarro JC, van Raaij MJ, Ramelot TA, Rousseau F, Tong
L, Wernimont AK, Young J, Schwede T (2011) Target highlights in
CASP9: Experimental target structures for the critical assessment
of techniques for protein structure prediction. Proteins 79 S10,
6-20.
- Leiman PG, Arisaka F, van Raaij MJ, Kostyuchenko VA, Aksyuk AA,
Kanamaru S, Rossmann MG (2010) Morphogenesis of the T4 tail and
tail fibers. Virol J 7, 355.
- Bartual SG, Otero JM, Garcia-Doval C, Llamas-Saiz AL, Kahn R,
Fox GC, van Raaij MJ (2010) Structure of the bacteriophage T4 long
tail fiber needle-shaped receptor-binding tip. PNAS USA 107,
20287-20292.
- Bartual SG, Garcia-Doval C, Alonso J, Schoehn G, van Raaij MJ
(2010) Two-chaperone assisted soluble expression and purification
of the bacteriophage T4 long tail fibre protein gp37. Prot Expr
Purif 70, 116-121.
- Walter M, Fiedler C, Grassl R, Biebl M, Hermo-Parrado XL,
Llamas-Saiz AL, Seckler R, Miller S, van Raaij MJ(2008) Structure
of the receptor-binding protein of bacteriophage Det7: a podoviral
tail spike in a myovirus. J
Virol. 82 2265-2273.
- Mitraki A,
Papanikolopoulou K, van Raaij MJ (2006) Natural triple beta-stranded fibrous folds. Adv Protein Chem 73 (Fibrous Proteins:
Amyloids, Prions and Beta Proteins), 97-124.
- Mitraki A, van Raaij MJ
(2005) Folding of beta-structured fibrous proteins and
self-assembling peptides. Meth. Mol Biol. 300 (Protein
Nanotechnology: Protocols, Instrumentation, and Application),
125-140.
- Papanikolopoulou K, Teixeira S, Belrhali H, Forsyth VT,
Mitraki A, van Raaij MJ (2004) Adenovirus fibre shaft sequences
fold into the native triple beta-spiral fold when N-terminally
fused to the bacteriophage T4 fibritin foldon trimerisation motif.
J Mol Biol 342, 219-227.
- van Raaij MJ, Mitraki A (2004) Beta-structured viral fibres:
assembly, structure and implications for materials design. Curr
Opin Solid State Mater Sci 8, 151-156.
- Kostyuchenko VA, Leiman PG, Chipman PR, Kanamaru S, van
Raaij MJ, Arisaka F, Mesyanzhinov VV, Rossmann MG (2003) Three-dimensional structure of bacteriophage T4
baseplate. Nat Struct Biol 10, 688-693.
- Thomassen E, Gielen G, Schuetz M, Schoehn G, Abrahams JP,
Miller S, van Raaij MJ (2003) The structure of the
receptor-binding domain of the bacteriophage T4 short tail fibre
reveals a knitted trimeric metal-binding fold. J Mol Biol
331, 361-373.
- Mitraki A, Miller S, van Raaij MJ (2002) Review: conformation
and folding of novel beta-structural elements in viral fiber
proteins: the triple beta-spiral and triple beta-helix. J
Struct Biol 137, 236-247.
- van Raaij MJ, Schoehn G, Burda MR, Miller S (2001) Crystal
structure of a heat- and protease-stablepart of the bacteriophage
T4 short tail fibre. J Mol Biol 314, 1137-1147.
- van Raaij MJ, Schoehn G, Jaquinod M, Ashman K, Burda MR, Miller
S (2001) Identification and crystallisation of a heat- and
protease-stable fragment of the bacteriophage T4 short tail fibre.
Biol Chem 382, 1049-1055.